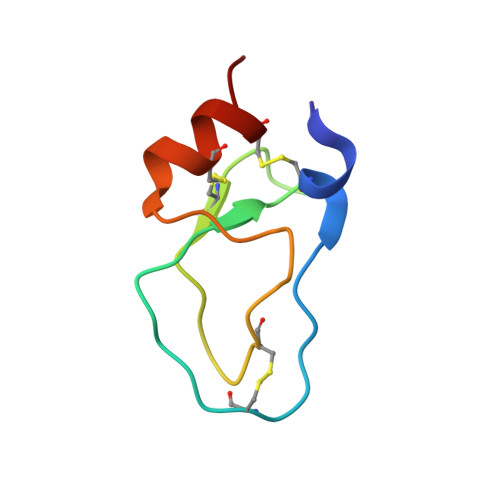
NMR structures of two variants of bovine pancreatic trypsin inhibitor (BPTI) reveal unexpected influence of mutations on protein structure and stability.
Cierpicki, T., Otlewski, J.(2002) J Mol Biology 321: 647-658
- PubMed: 12206780 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(02)00620-4
- Primary Citation Related Structures:
1LD5, 1LD6 - PubMed Abstract:
Here we determined NMR solution structures of two mutants of bovine pancreatic trypsin inhibitor (BPTI) to reveal structural reasons of their decreased thermodynamic stability. A point mutation, A16V, in the solvent-exposed loop destabilizes the protein by 20 degrees C, in contrast to marginal destabilization observed for G, S, R, L or W mutants. In the second mutant introduction of eight alanine residues at proteinase-contacting sites (residues 11, 13, 17, 18, 19, 34, 37 and 39) provides a protein that denatures at a temperature about 30 degrees C higher than expected from additive behavior of individual mutations. In order to efficiently determine structures of these variants, we applied a procedure that allows us to share data between regions unaffected by mutation(s). NOAH/DYANA and CNS programs were used for a rapid assignment of NOESY cross-peaks, structure calculations and refinement. The solution structure of the A16V mutant reveals no conformational change within the molecule, but shows close contacts between V16, I18 and G36/G37. Thus, the observed 4.3kcal/mol decrease of stability results from a strained local conformation of these residues caused by introduction of a beta-branched Val side-chain. Contrary to the A16V mutation, introduction of eight alanine residues produces significant conformational changes, manifested in over a 9A shift of the Y35 side-chain. This structural rearrangement provides about 6kcal/mol non-additive stabilization energy, compared to the mutant in which G37 and R39 are not mutated to alanine residues.
- Laboratory of Protein Engineering, Institute of Biochemistry and Molecular Biology, University of Wroclaw, Tamka 2, 50-137, Wroclaw, Poland.
Organizational Affiliation: