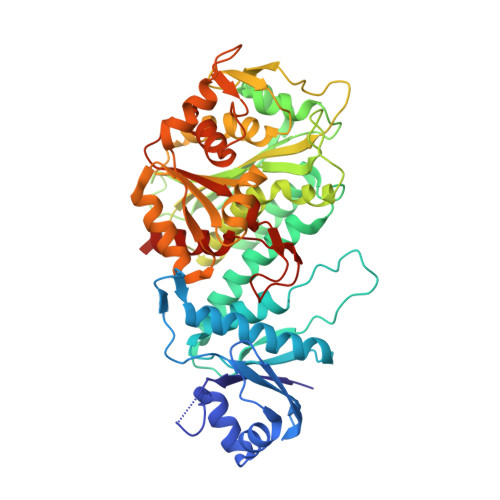
Molecular structure of leucine aminopeptidase at 2.7-A resolution.
Burley, S.K., David, P.R., Taylor, A., Lipscomb, W.N.(1990) Proc Natl Acad Sci U S A 87: 6878-6882
- PubMed: 2395881 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.87.17.6878
- Primary Citation Related Structures:
1LAP - PubMed Abstract:
The three-dimensional structure of bovine lens leucine aminopeptidase (EC 3.4.11.1) complexed with bestatin, a slow-binding inhibitor, has been solved to 3.0-A resolution by the multiple isomorphous replacement method with phase combination and density modification. In addition, the structure of the isomorphous native enzyme has been refined at 2.7-A resolution, and the current crystallographic R factor is 0.169 for a model that includes the two zinc ions and all 487 amino acid residues comprising the asymmetric unit. The enzyme is physiologically active as a hexamer, which has 32 symmetry and is triangular in shape with a triangle edge length of 115 A and maximal thickness of 90 A. The monomers are crystallographically equivalent and each is folded into two unequal alpha/beta domains connected by an alpha-helix to give a comma-like shape with approximate maximal dimensions of 90 x 55 x 55 A3. The secondary structural composition is 40% alpha-helix and 19% beta-strand. The N-terminal domain (160 amino acids) mediates trimer-trimer interactions and does not appear to participate directly in catalysis. The C-terminal domain (327 amino acids) is responsible for catalysis and binds the two zinc ions, which are 2.88 A apart. The pair of metal ions is located near the edge of an eight-stranded, saddle-shaped beta-sheet. One zinc ion is coordinated by carboxylate oxygen atoms of Asp-255, Asp-332, and Glu-334 and the carbonyl oxygen of Asp-332. The other zinc ion is coordinated by the carboxylate oxygen atoms of Asp-255, Asp-273, and Glu-334. The active site also contains two positively charged residues, Lys-250 and Arg-336. The six active sites are themselves located in the interior of the hexamer, where they line a disk-shaped cavity of radius 15 A and thickness 10 A. Access to this cavity is provided by solvent channels that run along the twofold symmetry axes.
- Gibbs Chemical Laboratory, Harvard University, Cambridge, MA 02138.
Organizational Affiliation: