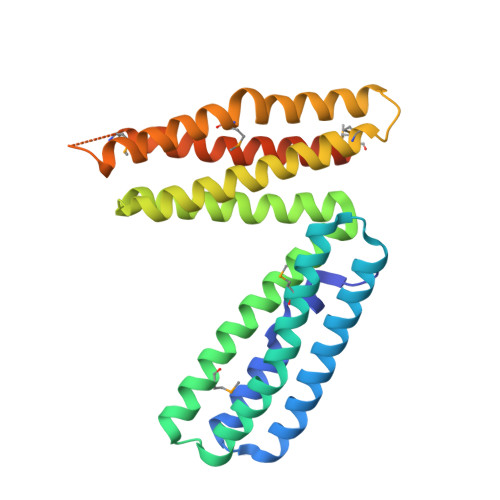
Biochemical and structural definition of the l-afadin- and actin-binding sites of alpha-catenin.
Pokutta, S., Drees, F., Takai, Y., Nelson, W.J., Weis, W.I.(2002) J Biological Chem 277: 18868-18874
- PubMed: 11907041 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M201463200
- Primary Citation Related Structures:
1L7C - PubMed Abstract:
alpha-Catenin is an integral component of adherens junctions, where it links cadherins to the actin cytoskeleton. alpha-Catenin is also required for the colocalization of the nectin/afadin/ponsin adhesion system to adherens junctions, and it specifically associates with the nectin-binding protein afadin. A proteolytic fragment of alpha-catenin, residues 385-651, contains the afadin-binding site. The three-dimensional structure of this fragment comprises two side-by-side four-helix bundles, both of which are required for afadin binding. The alpha-catenin fragment 385-651 binds afadin more strongly than the full-length protein, suggesting that the full-length protein harbors a cryptic binding site for afadin. Comparison of the alpha-catenin 385-651 structure with the recently solved structure of the alpha-catenin M-fragment (Yang, J., Dokurno, P., Tonks, N. K., and Barford, D. (2001) EMBO J. 20, 3645-3656) reveals a surprising flexibility in the orientation of the two four-helix bundles. alpha-Catenin and the actin-binding protein vinculin share sequence and most likely structural similarity within their actin-binding domains. Despite this homology, actin binding requires additional sequences adjacent to this region.
- Department of Structural Biology, Stanford University School of Medicine, Stanford, California 94305, USA.
Organizational Affiliation: