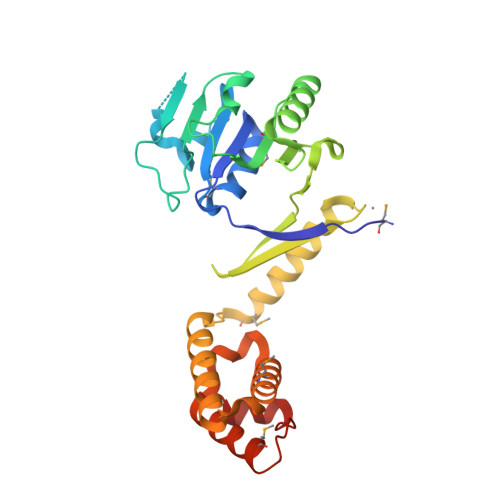
The structure of Saccharomyces cerevisiae Met8p, a bifunctional dehydrogenase and ferrochelatase.
Schubert, H.L., Raux, E., Brindley, A.A., Leech, H.K., Wilson, K.S., Hill, C.P., Warren, M.J.(2002) EMBO J 21: 2068-2075
- PubMed: 11980703 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/21.9.2068
- Primary Citation Related Structures:
1KYQ - PubMed Abstract:
Sirohaem is a tetrapyrrole-derived prosthetic group that is required for the essential assimilation of sulfur and nitrogen into all living systems as part of the sulfite and nitrite reductase systems. The final two steps in the biosynthesis of sirohaem involve a beta-NAD(+)-dependent dehydrogenation of precorrin-2 to generate sirohydrochlorin followed by ferrochelation to yield sirohaem. In Saccharomyces cerevisiae, Met8p is a bifunctional enzyme that carries out both of these reactions. Here, we report the 2.2 A resolution crystal structure of Met8p, which adopts a novel fold that bears no resemblance to the previously determined structures of cobalt- or ferro-chelatases. Analysis of mutant proteins suggests that both catalytic activities share a single active site, and that Asp141 plays an essential role in both dehydrogenase and chelatase processes.
- Department of Biochemistry, University of Utah, Salt Lake City, UT 84132, USA. heidi@biochem.utah.edu
Organizational Affiliation: