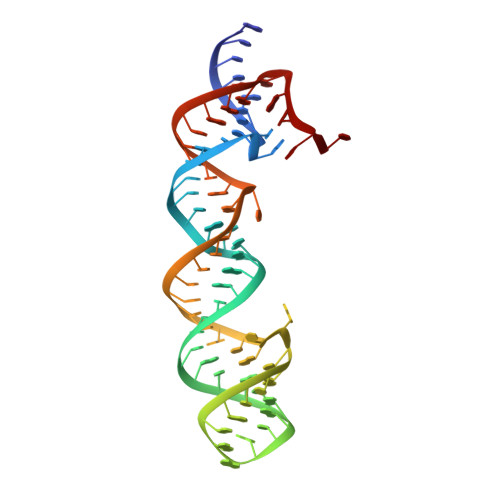
Structural insights into group II intron catalysis and branch-site selection.
Zhang, L., Doudna, J.A.(2002) Science 295: 2084-2088
- PubMed: 11859154 Search on PubMed
- DOI: https://doi.org/10.1126/science.1069268
- Primary Citation Related Structures:
1KXK - PubMed Abstract:
Group II self-splicing introns catalyze autoexcision from precursor RNA transcripts by a mechanism strikingly similar to that of the spliceosome, an RNA-protein assembly responsible for splicing together the protein-coding parts of most eukaryotic pre-mRNAs. Splicing in both cases initiates via nucleophilic attack at the 5' splice site by the 2' OH of a conserved intron adenosine residue, creating a branched (lariat) intermediate. Here, we describe the crystal structure at 3.0 A resolution of a 70-nucleotide RNA containing the catalytically essential domains 5 and 6 of the yeast ai5gamma group II self-splicing intron, revealing an unexpected two-nucleotide bulged structure around the branch-point adenosine in domain 6.
- Department of Molecular Biophysics and Biochemistry and, Howard Hughes Medical Institute, Yale University, New Haven, CT 06520, USA.
Organizational Affiliation: