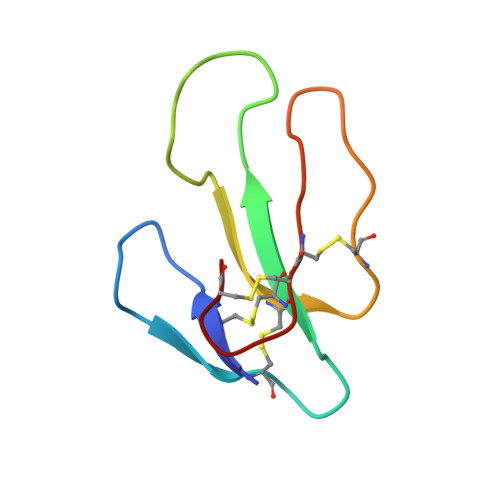
Crystal structure of cardiotoxin V from Taiwan cobra venom: pH-dependent conformational change and a novel membrane-binding motif identified in the three-finger loops of P-type cardiotoxin.
Sun, Y.J., Wu, W.G., Chiang, C.M., Hsin, A.Y., Hsiao, C.D.(1997) Biochemistry 36: 2403-2413
- PubMed: 9054545 Search on PubMed
- DOI: https://doi.org/10.1021/bi962594h
- Primary Citation Related Structures:
1KXI - PubMed Abstract:
The crystal structure of cardiotoxin V from Taiwan cobra venom (CTX A5) has been solved at pH 8.5 and refined to an R-factor of 20.7% for 7013 reflections [>2sigma(F)] between 8- and 2.19-A resolution. The refined model shows that CTX A5 exists as a dimer. The assembly consists of 974 non-hydrogen atoms from 124 residues and 73 water molecules. The global monomeric structure is similar to that determined by NMR at pH 3.7, characterized by a core formed by two beta-sheets connected with three-finger loops. However, local conformational differences are detected in two functionally important regions, loops I and II. A disparity between the NMR and X-ray structure of CTX A5 is detected near the tip of loop I and can be attributed to the difference in the protonation state of His4 at different pH, resulting in a reorientation of the His4 imidazole ring. A concerted motion of amino acid side chains located near His4 is detected and possibly contributes to the pH-dependent binding ability of CTX A5 to phospholipid model membranes. The second difference, detected at the tip of loop II, is due to the hydrophobic contact between CTX dimers in the crystal packing and the interaction of water molecules with amino acid residues in the loop II region of the CTX containing Pro31 (P-type CTX). This interaction forces loop II into a more rigid omega shape bridging the main chain at positions 27 and 34, contradictory to the flexible, tapering shape detected by NMR. Thus, a novel continuous hydrophobic column capable of binding to and possibly penetrating the membrane lipid bilayer is formed by the tips of the three-finger loops. In this respect, the X-ray crystal structure of CTX A5 may represent the CTX structure in the membrane-binding mode.
- Crystallography Laboratory, Institute of Molecular Biology, Academia Sinica, Taipei, Taiwan.
Organizational Affiliation: