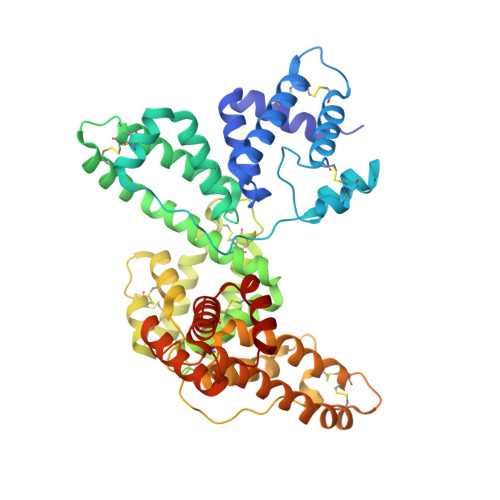
Crystal structures of the vitamin D-binding protein and its complex with actin: structural basis of the actin-scavenger system.
Otterbein, L.R., Cosio, C., Graceffa, P., Dominguez, R.(2002) Proc Natl Acad Sci U S A 99: 8003-8008
- PubMed: 12048248 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.122126299
- Primary Citation Related Structures:
1KW2, 1KXP - PubMed Abstract:
Actin is the most abundant protein in eukaryotic cells, but its release from cells into blood vessels can be lethal, being associated with clinical situations including hepatic necrosis and septic shock. A homeostatic mechanism, termed the actin-scavenger system, is responsible for the depolymerization and removal of actin from the circulation. During the first phase of this mechanism, gelsolin severs the actin filaments. In the second phase, the vitamin D-binding protein (DBP) traps the actin monomers, which accelerates their clearance. We have determined the crystal structures of DBP by itself and complexed with actin to 2.1 A resolution. Similar to its homologue serum albumin, DBP consists of three related domains. Yet, in DBP a strikingly different organization of the domains gives rise to a large actin-binding cavity. After complex formation the three domains of DBP move slightly to "clamp" onto actin subdomain 3 and to a lesser extent subdomain 1. Contacts between actin and DBP throughout their extensive 3,454-A(2) intermolecular interface involve a mixture of hydrophobic, electrostatic, and solvent-mediated interactions. The area of actin covered by DBP within the complex approximately equals the sum of those covered by gelsolin and profilin. Moreover, certain interactions of DBP with actin mirror those observed in the actin-gelsolin complex, which may explain how DBP can compete effectively with gelsolin for actin binding. Formation of the strong actin-DBP complex proceeds with limited conformational changes to both proteins, demonstrating how DBP has evolved to become an effective actin-scavenger protein.
- Boston Biomedical Research Institute, 64 Grove Street, Watertown, MA 02472, USA.
Organizational Affiliation: