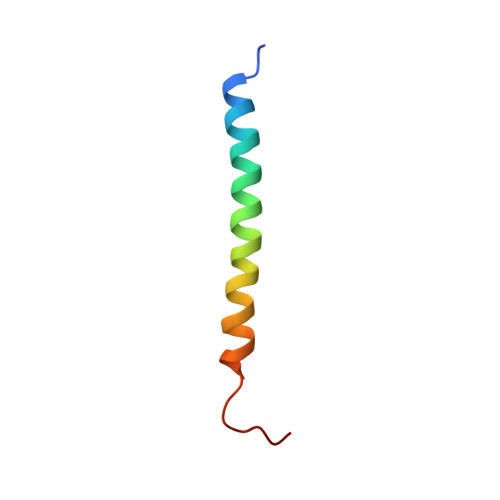
Solution structure of moricin, an antibacterial peptide, isolated from the silkworm Bombyx mori.
Hemmi, H., Ishibashi, J., Hara, S., Yamakawa, M.(2002) FEBS Lett 518: 33-38
- PubMed: 11997013 Search on PubMed
- DOI: https://doi.org/10.1016/s0014-5793(02)02637-6
- Primary Citation Related Structures:
1KV4 - PubMed Abstract:
A novel antibacterial peptide, moricin, isolated from the silkworm Bombyx mori, consists of 42 amino acids. It is highly basic and the amino acid sequence has no significant similarity to those of other antibacterial peptides. The 20 structures of moricin in methanol have been determined from two-dimensional 1H-nuclear magnetic resonance spectroscopic data. The solution structure reveals an unique structure comprising of a long alpha-helix containing eight turns along nearly the full length of the peptide except for four N-terminal residues and six C-terminal residues. The electrostatic surface map shows that the N-terminal segment of the alpha-helix, residues 5-22, is an amphipathic alpha-helix with a clear separation of hydrophobic and hydrophilic faces, and that the C-terminal segment of the alpha-helix, residues 23-36, is a hydrophobic alpha-helix except for the negatively charged surface at the position of Asp30. The results suggest that the amphipathic N-terminal segment of the alpha-helix is mainly responsible for the increase in permeability of the membrane to kill the bacteria.
- National Food Research Institute, 2-1-12 Kannondai, Tsukuba, Ibaraki 305-8642, Japan. hemmi@nfri.affrc.go.jp
Organizational Affiliation: