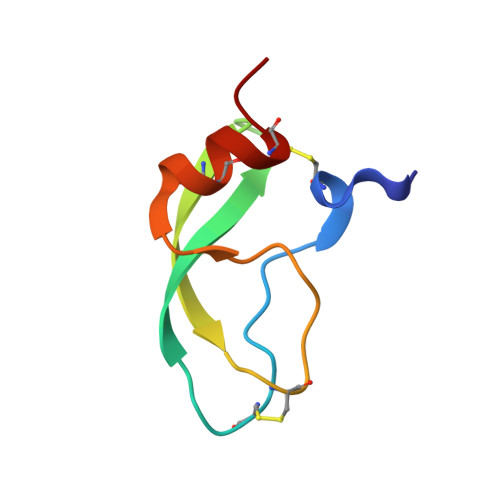
Solution structure and backbone dynamics of the human alpha3-chain type VI collagen C-terminal Kunitz domain,.
Sorensen, M.D., Bjorn, S., Norris, K., Olsen, O., Petersen, L., James, T.L., Led, J.J.(1997) Biochemistry 36: 10439-10450
- PubMed: 9265624 Search on PubMed
- DOI: https://doi.org/10.1021/bi9705570
- Primary Citation Related Structures:
1KUN - PubMed Abstract:
The solution structure and backbone dynamics of the 58-residue C-terminal Kunitz domain fragment [alpha3(VI)] of human alpha3-chain type VI collagen has been studied by two-dimensional 1H-1H and 1H-15N nuclear magnetic resonance spectroscopy at 303 K. The solution structure is represented by an ensemble of 20 structures calculated with X-PLOR using 612 distance and 47 dihedral angle restraints. The distance restraints were obtained by a complete relaxation matrix analysis using MARDIGRAS. The root mean squared (rms) deviation is 0.91 A for the backbone atoms of the residues Thr2(8)-Gly12(18), Arg15(21)-Tyr35(41), and Gly40(46)-Pro57(63). The central beta-sheet [residues Ile18(24)-Tyr35(41)] and the C-terminal alpha-helix [residues Gln48(54)-Cys55(61)] are better defined with a backbone rms deviation of 0.46 A. The solution structure of alpha3(VI) is virtually identical to the crystal structure of alpha3(VI) and to the solution structure of bovine pancreatic trypsin inhibitor (BPTI). The 15N spin-lattice and spin-spin relaxation rates and the 1H-15N heteronuclear nuclear Overhauser enhancement (NOE) were analyzed using both the "model-free" formalism [Lipari, G., & Szabo, A. (1982) J. Am. Chem. Soc. 104, 4546-4559 and 4559-4570] and the reduced spectral density mapping procedure [Farrow, N. A., Szabo, A., Torchia, D. A., & Kay, L. E. (1995) J. Biomol.NMR 6, 153-162]. The results obtained from the "model-free" analysis include an overall correlation time tauc of 3. 00 ns and backbone order parameters S2 in the range from 0.28 to 0. 93. The necessity of including an exchange term in the analysis of the relaxation data from 14 residues indicated that these residues are involved in motions on the micro- to millisecond time scale. The majority of the 14 residues are located in the vicinity of the Cys14(20)-Cys38(44) disulfide bond, suggesting the presence of a disulfide bond isomerization similar to the one observed in BPTI [Otting, G., Liepinsh, E., & Wüthrich, K. (1993) Biochemistry 32, 3571-3582]. It is suggested that this disulfide bond isomerization is the main reason for the surprisingly small effect on trypsin inhibition observed when Thr13(19) of alpha3(VI) is substituted with Pro.
- Department of Chemistry, University of Copenhagen, The H. C. Orsted Institute, Universitetsparken 5, DK-2100 Copenhagen O, Denmark.
Organizational Affiliation: