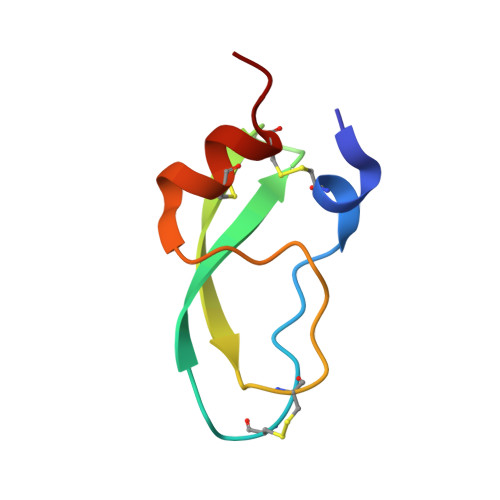
Anisotropic behaviour of the C-terminal Kunitz-type domain of the alpha3 chain of human type VI collagen at atomic resolution (0.9 A).
Arnoux, B., Ducruix, A., Prange, T.(2002) Acta Crystallogr D Biol Crystallogr 58: 1252-1254
- PubMed: 12077460 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444902007333
- Primary Citation Related Structures:
1KTH - PubMed Abstract:
The C-terminal Kunitz-type domain from the alpha3 chain of human type VI collagen (C5), a single amino-acid residue chain with three disulfide bridges, was refined at 0.9 A resolution in a monoclinic form, space group P2(1) with one molecule per asymmetric unit, using data collected at cryogenic temperature (110 K). The average protein factor decreases from 21 A(2) at room temperature (RT) to 12 A(2) at cryotemperature (100 K, CT). The spatially close N- and C-termini remain highly disordered. The different structural motifs of C5 were analyzed in terms of rigid-body displacement (TLS analyses) and show dominant libration motion for the secondary structure.
- Laboratoire de Cristallographie et RMN Biologiques (UMR 8015 CNRS), Faculté de Pharmacie 4, Avenue de l'Observatoire, 75006 Paris, France. arnoux@pharmacie.univ-paris5.fr
Organizational Affiliation: