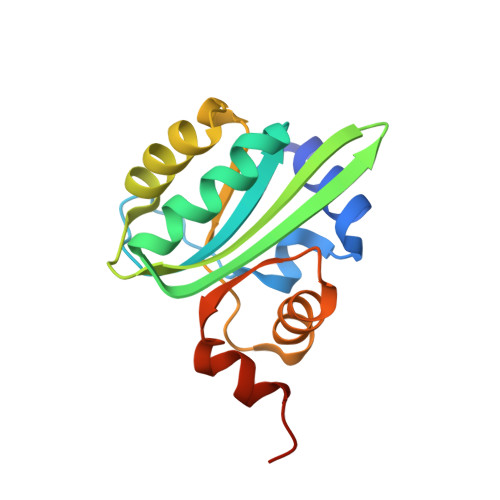
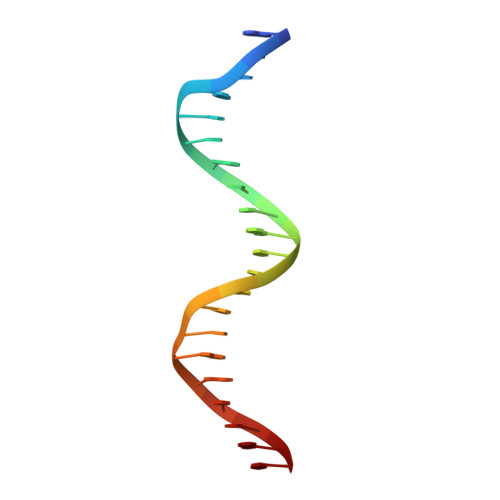
Crystal structures of two intermediates in the assembly of the papillomavirus replication initiation complex.
Enemark, E.J., Stenlund, A., Joshua-Tor, L.(2002) EMBO J 21: 1487-1496
- PubMed: 11889054 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/21.6.1487
- Primary Citation Related Structures:
1KSX, 1KSY - PubMed Abstract:
Initiation of DNA replication of the papillomavirus genome is a multi-step process involving the sequential loading of viral E1 protein subunits onto the origin of replication. Here we have captured structural snapshots of two sequential steps in the assembly process. Initially, an E1 dimer binds to adjacent major grooves on one face of the double helix; a second dimer then binds to another face of the helix. Each E1 monomer has two DNA-binding modules: a DNA-binding loop, which binds to one DNA strand and a DNA-binding helix, which binds to the opposite strand. The nature of DNA binding suggests a mechanism for the transition between double- and single-stranded DNA binding that is implicit in the progression to a functional helicase.
- W.M.Keck Structural Biology Laboratory, 1 Bungtown Road, Cold Spring Harbor, NY 11724, USA.
Organizational Affiliation: