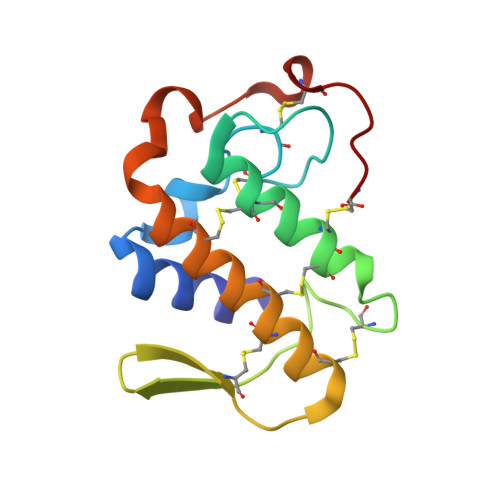
First Structural Evidence of a Specific Inhibition of Phospholipase A2 by alpha-Tocopherol (Vitamin E) and its Implications in Inflammation: Crystal Structure of the Complex Formed Between Phospholipase A2 and alpha-Tocopherol at 1.8 A Resolution
Chandra, V., Jasti, J., Kaur, P., Betzel, C., Srinivasan, A., Singh, T.P.(2002) J Mol Biology 320: 215-222
- PubMed: 12079380 Search on PubMed
- DOI: https://doi.org/10.1016/S0022-2836(02)00473-4
- Primary Citation Related Structures:
1KPM - PubMed Abstract:
This is the first structural evidence of alpha-tocopherol (alpha-TP) as a possible candidate against inflammation, as it inhibits phospholipase A2 specifically and effectively. The crystal structure of the complex formed between Vipera russelli phospholipase A2 and alpha-tocopherol has been determined and refined to a resolution of 1.8 A. The structure contains two molecules, A and B, of phospholipase A2 in the asymmetric unit, together with one alpha-tocopherol molecule, which is bound specifically to one of them. The phospholipase A2 molecules interact extensively with each other in the crystalline state. The two molecules were found in a stable association in the solution state as well, thus indicating their inherent tendency to remain together as a structural unit, leading to significant functional implications. In the crystal structure, the most important difference between the conformations of two molecules as a result of their association pertains to the orientation of Trp31. It may be noted that Trp31 is located at the mouth of the hydrophobic channel that forms the binding domain of the enzyme. The values of torsion angles (phi, psi, chi(1) and chi(2)) for both the backbone as well as for the side-chain of Trp31 in molecules A and B are -94 degrees, -30 degrees, -66 degrees, 116 degrees and -128 degrees, 170 degrees, -63 degrees, -81 degrees, respectively. The conformation of Trp31 in molecule A is suitable for binding, while that in B hinders the passage of the ligand to the binding site. Consequently, alpha-tocopherol is able to bind to molecule A only, while the binding site of molecule B contains three water molecules. In the complex, the aromatic moiety of alpha-tocopherol is placed in the large space at the active site of the enzyme, while the long hydrophobic channel in the enzyme is filled by hydrocarbon chain of alpha-tocopherol. The critical interactions between the enzyme and alpha-tocopherol are generated between the hydroxyl group of the six-membered ring of alpha-tocopherol and His48 N(delta1) and Asp49 O(delta1) as characteristic hydrogen bonds. The remaining part of alpha-tocopherol interacts extensively with the residues of the hydrophobic channel of the enzyme, giving rise to a number of hydrophobic interactions, resulting in the formation of a stable complex.
- Department of Biophysics, All India Institute of Medical Sciences, New Delhi 110029, India.
Organizational Affiliation: