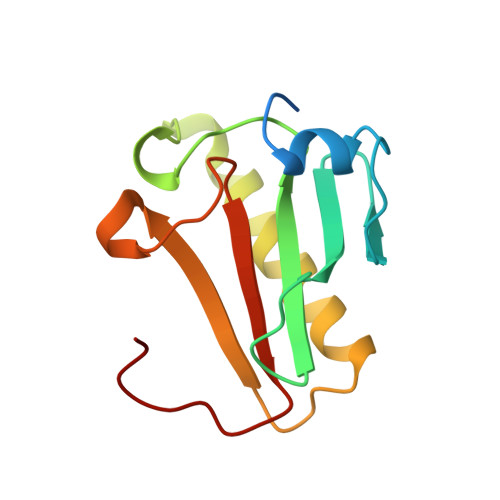
Three-dimensional structure of human protein kinase C interacting protein 1, a member of the HIT family of proteins.
Lima, C.D., Klein, M.G., Weinstein, I.B., Hendrickson, W.A.(1996) Proc Natl Acad Sci U S A 93: 5357-5362
- PubMed: 8643579 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.93.11.5357
- Primary Citation Related Structures:
1KPA, 1KPB, 1KPC - PubMed Abstract:
The three-dimensional structure of protein kinase C interacting protein 1 (PKCI-1) has been solved to high resolution by x-ray crystallography using single isomorphous replacement with anomalous scattering. The gene encoding human PKCI-1 was cloned from a cDNA library by using a partial sequence obtained from interactions identified in the yeast two-hybrid system between PKCI-1 and the regulatory domain of protein kinase C-beta. The PKCI-1 protein was expressed in Pichia pastoris as a dimer of two 13.7-kDa polypeptides. PKCI-1 is a member of the HIT family of proteins, shown by sequence identity to be conserved in a broad range of organisms including mycoplasma, plants, and humans. Despite the ubiquity of this protein sequence in nature, no distinct function has been shown for the protein product in vitro or in vivo. The PKCI-1 protomer has an alpha+beta meander fold containing a five-stranded antiparallel sheet and two helices. Two protomers come together to form a 10-stranded antiparallel sheet with extensive contacts between a helix and carboxy terminal amino acids of a protomer with the corresponding amino acids in the other protomer. PKCI-1 has been shown to interact specifically with zinc. The three-dimensional structure has been solved in the presence and absence of zinc and in two crystal forms. The structure of human PKCI-1 provides a model of this family of proteins which suggests a stable fold conserved throughout nature.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032, USA.
Organizational Affiliation: