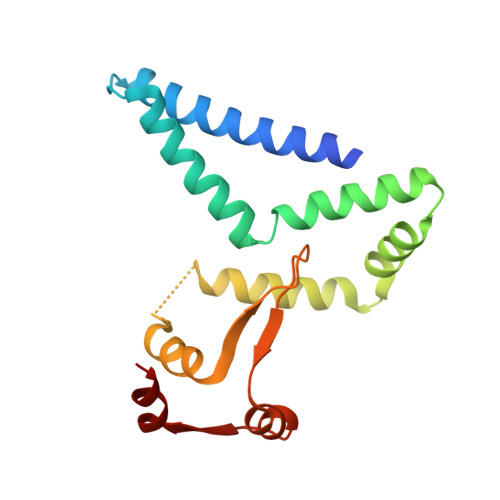
Recognition of the rotavirus mRNA 3' consensus by an asymmetric NSP3 homodimer.
Deo, R.C., Groft, C.M., Rajashankar, K.R., Burley, S.K.(2002) Cell 108: 71-81
- PubMed: 11792322 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(01)00632-8
- Primary Citation Related Structures:
1KNZ - PubMed Abstract:
Rotaviruses, the cause of life-threatening diarrhea in humans and cattle, utilize a functional homolog of poly(A) binding protein (PABP) known as nonstructural protein 3 (NSP3) for translation of viral mRNAs. NSP3 binds to viral mRNA 3' consensus sequences and circularizes the mRNA via interactions with eIF4G. The X-ray structure of the NSP3 RNA binding domain bound to a rotaviral mRNA 3' end has been determined. NSP3 is a novel, heart-shaped homodimer with a medial RNA binding cleft. The homodimer is asymmetric, and contains two similar N-terminal segments plus two structurally different C-terminal segments that intertwine to create a tunnel enveloping the mRNA 3' end. Biophysical studies demonstrate high affinity binding leading to increased thermal stability and slow dissociation kinetics, consistent with NSP3 function.
- Laboratories of Molecular Biophysics, The Rockefeller University, 1230 York Avenue, New York, NY 10021, USA.
Organizational Affiliation: