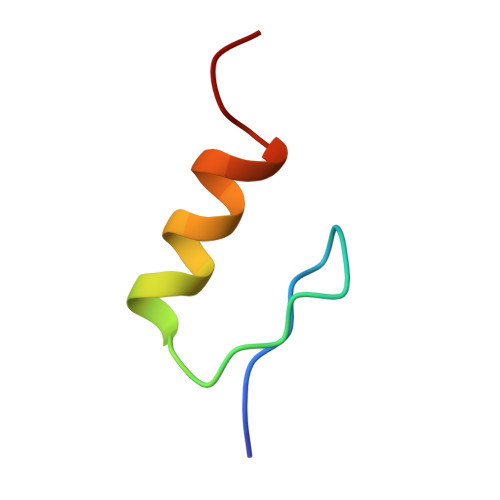
The hidden thermodynamics of a zinc finger.
Lachenmann, M.J., Ladbury, J.E., Phillips, N.B., Narayana, N., Qian, X., Weiss, M.A.(2002) J Mol Biology 316: 969-989
- PubMed: 11884136 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.5335
- Primary Citation Related Structures:
1KLR, 1KLS - PubMed Abstract:
The Zn finger provides a model for studies of protein structure and stability. Its core contains a conserved phenylalanine residue adjoining three architectural elements: a beta-hairpin, an alpha-helix and a tetrahedral Zn(2+)-binding site. Here, we demonstrate that the consensus Phe is not required for high-affinity Zn(2+) binding but contributes to the specification of a precise DNA-binding surface. Substitution of Phe by leucine in a ZFY peptide permits Zn(2+)-dependent folding. Although a native-like structure is retained, structural fluctuations lead to attenuation of selected nuclear Overhauser enhancements and accelerated amide proton exchange. Surprisingly, wild-type Zn affinity is maintained by entropy-enthalpy compensation (EEC): a hidden entropy penalty (TDeltaDeltaS 7kcal/mol) is balanced by enhanced enthalpy of association (DeltaDeltaH -7kcal/mol) at 25 degrees C. Because the variant is less well ordered than the Phe-anchored domain, the net change in entropy is opposite to the apparent change in configurational entropy. By analogy to the thermodynamics of organometallic complexation, we propose that EEC arises from differences in solvent reorganization. Exclusion of Leu among biological sequences suggests an evolutionary constraint on the dynamics of a Zn finger.
- Department of Biochemistry, Case Western Reserve University, 10900 Euclid Avenue, Cleveland, OH 44106-4935, USA.
Organizational Affiliation: