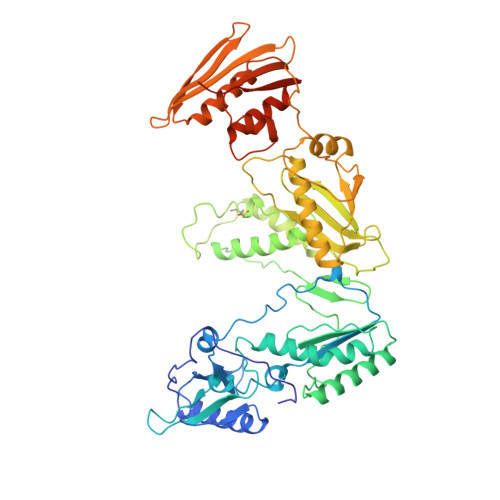
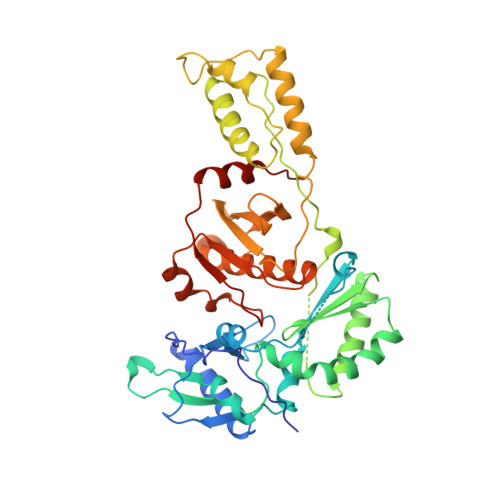
Unique features in the structure of the complex between HIV-1 reverse transcriptase and the bis(heteroaryl)piperazine (BHAP) U-90152 explain resistance mutations for this nonnucleoside inhibitor.
Esnouf, R.M., Ren, J., Hopkins, A.L., Ross, C.K., Jones, E.Y., Stammers, D.K., Stuart, D.I.(1997) Proc Natl Acad Sci U S A 94: 3984-3989
- PubMed: 9108091 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.94.8.3984
- Primary Citation Related Structures:
1KLM - PubMed Abstract:
The viral reverse transcriptase (RT) provides an attractive target in the search for anti-HIV therapies. The nonnucleoside inhibitors (NNIs) are a diverse set of compounds (usually HIV-1 specific) that function by distorting the polymerase active site upon binding in a nearby pocket. Despite being potent and of generally low toxicity, their clinical use has been limited by rapid selection for resistant viral populations. The 2.65-A resolution structure of the complex between HIV-1 RT and the bis(heteroaryl)piperazine (BHAP) NNI, 1-(5-methanesulfonamido-1H-indol-2-yl-carbonyl)-4- [3-(1-methyl-ethylamino) pyridinyl] piperazine (U-90152), reveals the inhibitor conformation and bound water molecules. The bulky U-90152 molecule occupies the same pocket as other NNIs, but the complex is stabilized quite differently, in particular by hydrogen bonding to the main chain of Lys-103 and extensive hydrophobic contacts with Pro-236. These interactions rationalize observed resistance mutations, notably Pro-236-Leu, which occurs characteristically for BHAPs. When bound, part of U-90152 protrudes into the solvent creating a channel between Pro-236 and the polypeptide segments 225-226 and 105-106, giving the first clear evidence of the entry mode for NNIs. The structure allows prediction of binding modes for related inhibitors [(altrylamino)piperidine-BHAPs] and suggests changes to U-90152, such as the addition of a 6 amino group to the pyridine ring, which may make binding more resilient to mutations in the RT. The observation of novel hydrogen bonding to the protein main chain may provide lessons for the improvement of quite different inhibitors.
- Laboratory of Molecular Biophysics, Oxford, United Kingdom.
Organizational Affiliation: