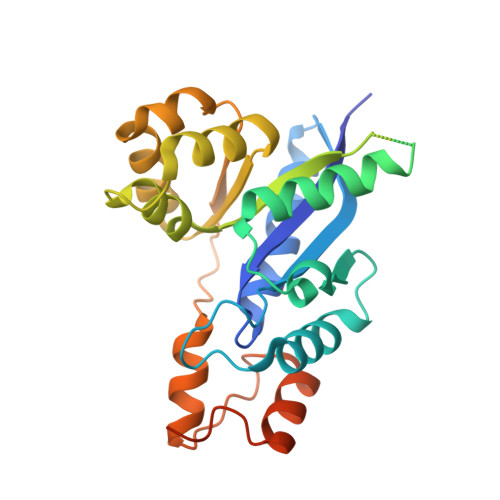
Structure of human NMN adenylyltransferase. A key nuclear enzyme for NAD homeostasis.
Garavaglia, S., D'Angelo, I., Emanuelli, M., Carnevali, F., Pierella, F., Magni, G., Rizzi, M.(2002) J Biological Chem 277: 8524-8530
- PubMed: 11751893 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M111589200
- Primary Citation Related Structures:
1KKU - PubMed Abstract:
Nicotinamide mononucleotide adenylyltransferase (NMNAT), a member of the nucleotidyltransferase alpha/beta-phosphodiesterases superfamily, catalyzes a universal step (NMN + ATP = NAD + PP(i)) in NAD biosynthesis. Localized within the nucleus, the activity of the human enzyme is greatly altered in tumor cells, rendering it a promising target for cancer chemotherapy. By using a combination of single isomorphous replacement and density modification techniques, the human NMNAT structure was solved by x-ray crystallography to a 2.5-A resolution, revealing a hexamer that is composed of alpha/beta-topology subunits. The active site topology of the enzyme, analyzed through homology modeling and structural comparison with other NMNATs, yielded convincing evidence for a substrate-induced conformational change. We also observed remarkable structural conservation in the ATP-recognition motifs GXXXPX(T/H)XXH and SXTXXR, which we take to be the universal signature for NMNATs. Structural comparison of human and prokaryotic NMNATs may also lead to the rational design of highly selective antimicrobial drugs.
- Department of Genetics and Microbiology A. Buzzati Traverso, University of Pavia, Via Ferrata 1, 27100 Pavia, Italy.
Organizational Affiliation: