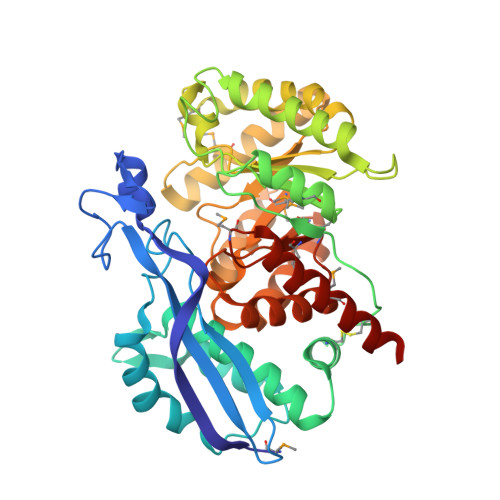
Insights into enzyme evolution revealed by the structure of methylaspartate ammonia lyase.
Levy, C.W., Buckley, P.A., Sedelnikova, S., Kato, Y., Asano, Y., Rice, D.W., Baker, P.J.(2002) Structure 10: 105-113
- PubMed: 11796115 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00696-7
- Primary Citation Related Structures:
1KKO, 1KKR - PubMed Abstract:
Methylaspartate ammonia lyase (MAL) catalyzes the magnesium-dependent reversible alpha,beta-elimination of ammonia from L-threo-(2S,3S)-3-methylaspartic acid to mesaconic acid. The 1.3 A MAD crystal structure of the dimeric Citrobacter amalonaticus MAL shows that each subunit comprises two domains, one of which adopts the classical TIM barrel fold, with the active site at the C-terminal end of the barrel. Despite very low sequence similarity, the structure of MAL is closely related to those of representative members of the enolase superfamily, indicating that the mechanism of MAL involves the initial abstraction of a proton alpha to the 3-carboxyl of (2S,3S)-3-methylasparic acid to yield an enolic intermediate. This analysis resolves the conflict that had linked MAL to the histidine and phenylalanine ammonia lyase family of enzymes.
- Krebs Institute for Biomolecular Research, Department of Molecular Biology and Biotechnology, The University of Sheffield, Sheffield S10 2TN, United Kingdom.
Organizational Affiliation: