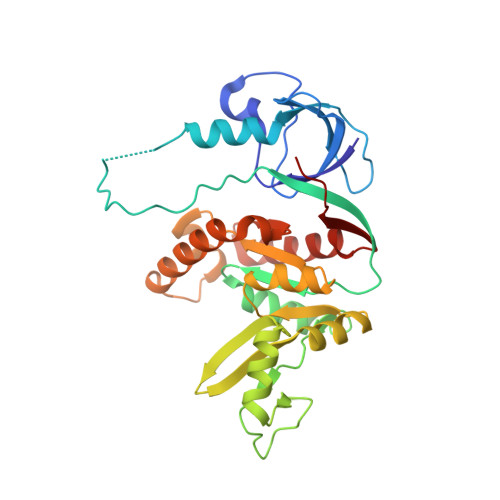
Structure of the SH3-Guanylate Kinase Module from PSD-95 Suggests a Mechanism for Regulated Assembly of MAGUK Scaffolding Proteins
McGee, A.W., Dakoji, S.R., Olsen, O., Bredt, D.S., Lim, W.A., Prehoda, K.E.(2001) Mol Cell 8: 1291-1301
- PubMed: 11779504 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(01)00411-7
- Primary Citation Related Structures:
1KJW - PubMed Abstract:
Membrane-associated guanylate kinases (MAGUKs), such as PSD-95, are modular scaffolds that organize signaling complexes at synapses and other cell junctions. MAGUKs contain PDZ domains, which recruit signaling proteins, as well as a Src homology 3 (SH3) and a guanylate kinase-like (GK) domain, implicated in scaffold oligomerization. The crystal structure of the SH3-GK module from PSD-95 reveals that these domains form an integrated unit: the SH3 fold comprises noncontiguous sequence elements divided by a hinge region and the GK domain. These elements compose two subdomains that can assemble in either an intra- or intermolecular fashion to complete the SH3 fold. We propose a model for MAGUK oligomerization in which complementary SH3 subdomains associate by 3D domain swapping. This model provides a possible mechanism for ligand regulation of oligomerization.
- Department of Physiology, University of California-San Francisco, San Francisco, CA 94143, USA.
Organizational Affiliation: