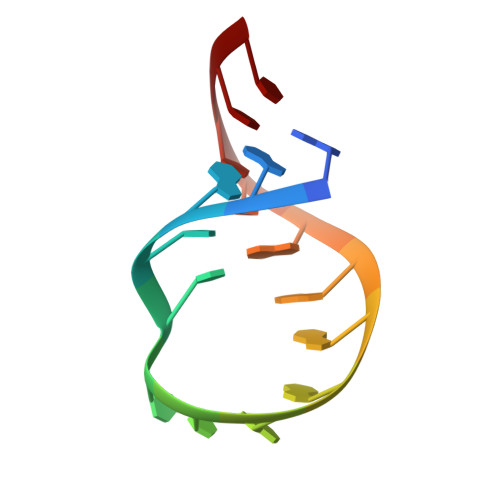
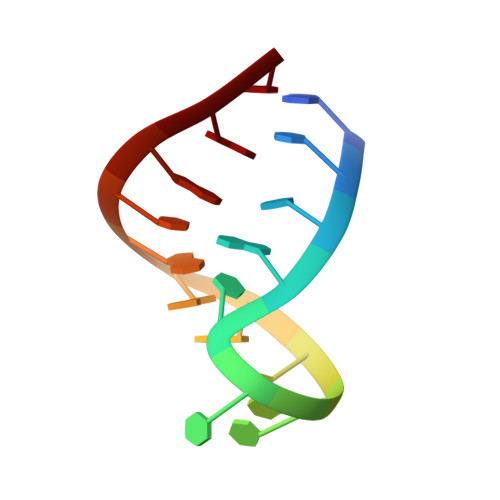
The structure of an RNA "kissing" hairpin complex of the HIV TAR hairpin loop and its complement.
Chang, K.Y., Tinoco Jr., I.(1997) J Mol Biology 269: 52-66
- PubMed: 9193000 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1021
- Primary Citation Related Structures:
1KIS - PubMed Abstract:
We have used nuclear magnetic resonance (NMR) to obtain the structure of an RNA "kissing" hairpin complex formed between the HIV-2 TAR hairpin loop and a hairpin with a complementary loop sequence. Kissing hairpins are important in natural antisense reactions; their complex is a specific target for protein binding. The complex has all six nucleotides of each loop paired to form a bent quasicontinuous helix of three coaxially stacked helices: two stems plus a loop-loop interaction helix. Experimental constraints derived from heteronuclear and homonuclear NMR data on 13C and 15N-labeled RNA led to a structure for the loop-loop helix with an average root-mean-square deviation of 0.83 (+/-0.10) A for 33 converged structures relative to the average structure. The loop-loop helix of the kissing complex is distorted compared to A-form RNA. Its major groove is blocked by the phosphodiester bonds that connect the first loop residue of each hairpin with its own stem, and it is flanked by two negatively charged phosphate clusters. The loop-loop helix has alternating helical twists between adjacent base-pairs. The base-pairs at the helix junctions are overwound and three base-pairs near the helix junctions adopt high propeller twists. All these changes reduce the distance needed for the bridging phosphodiester bonds connecting each stem and loop to cross the major groove of the loop-loop helix, and result in a deformed RNA helix with localized perturbations in the minor groove surface. The alternating helical twist pattern, plus other distortions in the loop-loop helix may be important for Rom protein recognition of the kissing hairpin complex.
- Department of Chemistry, University of California at Berkeley, 94720-1460, USA.
Organizational Affiliation: