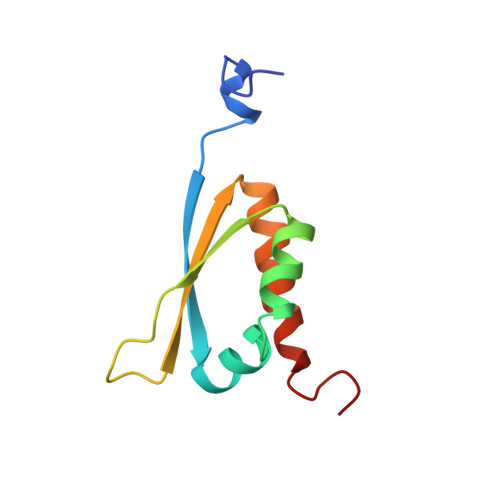
High precision solution structure of the C-terminal KH domain of heterogeneous nuclear ribonucleoprotein K, a c-myc transcription factor.
Baber, J.L., Libutti, D., Levens, D., Tjandra, N.(1999) J Mol Biology 289: 949-962
- PubMed: 10369774 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2818
- Primary Citation Related Structures:
1KHM - PubMed Abstract:
Among it's many reported functions, heterogeneous nuclear ribonucleoprotein (hnRNP) K is a transcription factor for the c- myc gene, a proto-oncogene critical for the regulation of cell growth and differentiation. We have determined the solution structure of the Gly26-->Arg mutant of the C-terminal K-homology (KH) domain of hnRNP K by NMR spectroscopy. This is the first structure investigation of hnRNP K. Backbone residual dipolar couplings, which provide information that is fundamentally different from the standard NOE-derived distance restraints, were employed to improve structure quality. An independent assessment of structure quality was achieved by comparing the backbone15N T1/T2ratios to the calculated structures. The C-terminal KH module of hnRNP K (KH3) is revealed to be a three-stranded beta-sheet stacked against three alpha-helices, two of which are nearly parallel to the strands of the beta-sheet. The Gly26-->Arg mutation abolishes single-stranded DNA binding without altering the overall fold of the protein. This provides a clue to possible nucleotide binding sites of KH3. It appears unlikely that the solvent-exposed side of the beta-sheet will be the site of protein-nucleic acid complex formation. This is in contrast to the earlier theme for protein-RNA complexes incorporating proteins structurally similar to KH3. We propose that the surface of KH3 that interacts with nucleic acid is comparable to the region of DNA interaction for the double-stranded DNA-binding domain of bovine papillomavirus-1 E2 that has a three-dimensional fold similar to that of KH3.
- National Heart, Lung, and Blood Institute, National Institutes of Health, Building 3, Bethesda, MD, 20892-0380, USA.
Organizational Affiliation: