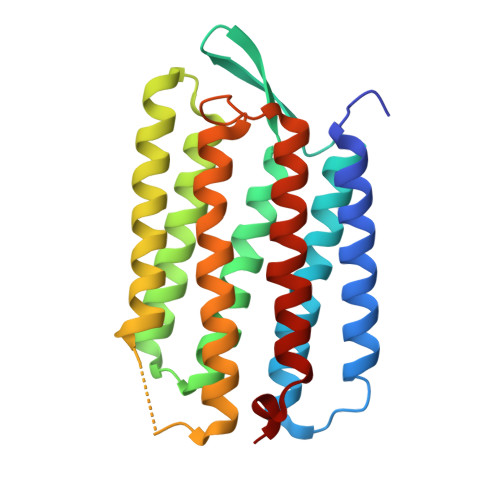
Structure of an early intermediate in the M-state phase of the bacteriorhodopsin photocycle.
Facciotti, M.T., Rouhani, S., Burkard, F.T., Betancourt, F.M., Downing, K.H., Rose, R.B., McDermott, G., Glaeser, R.M.(2001) Biophys J 81: 3442-3455
- PubMed: 11721006 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/S0006-3495(01)75976-0
- Primary Citation Related Structures:
1KG8, 1KG9, 1KGB - PubMed Abstract:
The structure of an early M-intermediate of the wild-type bacteriorhodopsin photocycle formed by actinic illumination at 230 K has been determined by x-ray crystallography to a resolution of 2.0 A. Three-dimensional crystals were trapped by illuminating with actinic light at 230 K, followed by quenching in liquid nitrogen. Amide I, amide II, and other infrared absorption bands, recorded from single bacteriorhodopsin crystals, confirm that the M-substate formed represents a structure that occurs early after deprotonation of the Schiff base. Rotation about the retinal C13-C14 double bond appears to be complete, but a relatively large torsion angle of 26 degrees is still seen for the C14-C15 bond. The intramolecular stress associated with the isomerization of retinal and the subsequent deprotonation of the Schiff base generates numerous small but experimentally measurable structural changes within the protein. Many of the residues that are displaced during the formation of the late M (M(N)) substate formed by three-dimensional crystals of the D96N mutant (Luecke et al., 1999b) are positioned, in early M, between their resting-state locations and the ones which they will adopt at the end of the M phase. The relatively small magnitude of atomic displacements observed in this intermediate, and the well-defined positions adopted by nearly all of the atoms in the structure, may make the formation of this structure favorable to model (simulate) by molecular dynamics.
- Graduate Group in Biophysics, Lawrence Berkeley National Laboratory, University of California, Berkeley, California 94720, USA.
Organizational Affiliation: