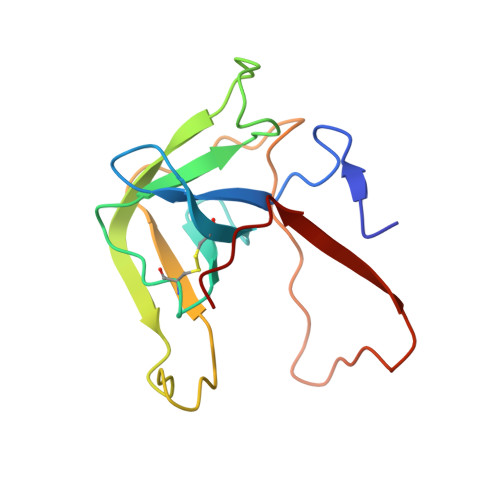
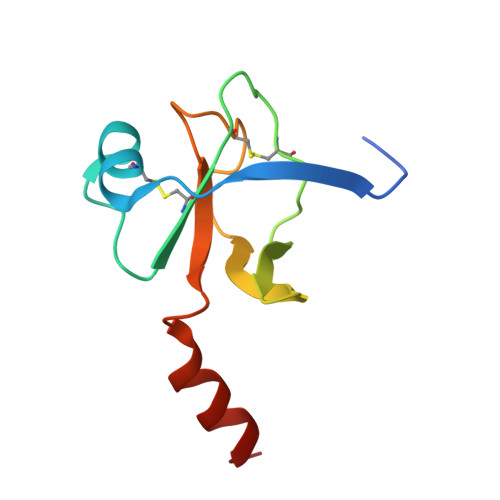
Three Dimensional Structures of S189D Chymotrypsin and D189S Trypsin Mutants: The Effect of Polarity at Site 189 on a Protease-specific Stabilization of the Substrate-binding Site
Szabo, E., Venekei, I., Bocskei, Z., Naray-Szabo, G., Graf, L.(2003) J Mol Biology 331: 1121-1130
- PubMed: 12927546 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00849-0
- Primary Citation Related Structures:
1KDQ - PubMed Abstract:
The crystal structure of S189D rat chymotrypsin have been determined (resolution 2.55A) and compared, together with D189S rat trypsin to wild-type structures to examine why these single mutations resulted in poorly active, non-specific enzymes instead of converting the specificities of trypsin and chymotrypsin into each other. Both mutants have stable structure but suffer from a surprisingly large number of serious deformations. These are restricted to the activation domain, mainly to the substrate-binding region and are larger in S189D chymotrypsin. A wild-type substrate-binding mode in the mutants is disfavored by substantial displacements of the Cys191-Cys220 disulfide and loop segments 185-195 (loop C2/D2) and 217-224 (loop E2/F2) at the specificity site. As a consequence, the substrate-binding clefts become wider and more solvent-accessible in the middle third and occluded in the lower third. Interestingly, while the Ser189 residue in D189S trypsin adopts a chymotrypsin-like conformation, the Asp189 residue in S189D chymotrypsin is turned out toward the solvent. The rearrangements in D189S trypsin are at the same sites where trypsin and trypsinogen differ and, in S189D chymotrypsin, the oxyanion hole as well as the salt-bridge between Asp194 and the N-terminal of Ile16 are missing as in chymotrypsinogen. Despite these similarities, the mutants do not have zymogen conformation. The Ser189Asp and Asp189Ser substitutions are structurally so disruptive probably because the stabilization of such a different specificity site polarities as those after the removal or introduction of a charged residue are beyond the capability of the wild-type conformation of the substrate-binding region.
- Department of Biochemistry, Hungarian Academy of Sciences, Eötvös Loránd University, 1117, Pázmány Péter sétány 1/A, Budapest, Hungary.
Organizational Affiliation: