The X-ray structure of human serum ceruloplasmin at 3.1 angstrom: Nature of the copper centres.
Zaitseva, I., Zaitsev, V., Card, G., Moshkov, K., Bax, B., Ralph, A., Lindley, P.(1996) J Biol Inorg Chem 1: 15-23
Experimental Data Snapshot
(1996) J Biol Inorg Chem 1: 15-23
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| CERULOPLASMIN | 1,046 | Homo sapiens | Mutation(s): 0 EC: 1.16.3.1 (PDB Primary Data), 1.16.3.4 (UniProt), 1.11.1.27 (UniProt), 1.11.1.9 (UniProt) | 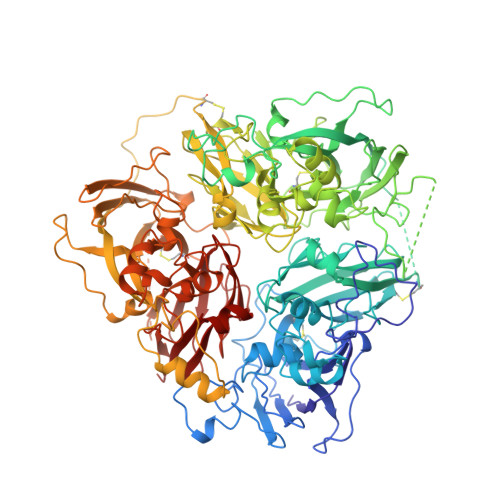 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P00450 GTEx: ENSG00000047457 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P00450 | ||||
Glycosylation | |||||
| Glycosylation Sites: 2 | Go to GlyGen: P00450-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Download:Ideal Coordinates CCD File | B [auth A], C [auth A] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| CU Download:Ideal Coordinates CCD File | D [auth A] E [auth A] F [auth A] G [auth A] H [auth A] | COPPER (II) ION Cu JPVYNHNXODAKFH-UHFFFAOYSA-N |  | ||
| O Download:Ideal Coordinates CCD File | L [auth A], M [auth A] | OXYGEN ATOM O XLYOFNOQVPJJNP-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 213.92 | α = 90 |
| b = 213.92 | β = 90 |
| c = 85.63 | γ = 120 |
| Software Name | Purpose |
|---|---|
| MOSFLM | data reduction |
| ROTAVATA | data reduction |
| X-PLOR | model building |
| X-PLOR | refinement |
| CCP4 | data scaling |
| X-PLOR | phasing |