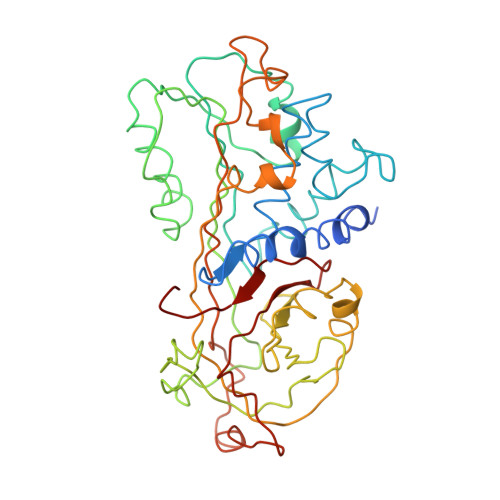
Crystal structure of an uncleaved alpha 1-antitrypsin reveals the conformation of its inhibitory reactive loop.
Song, H.K., Lee, K.N., Kwon, K.S., Yu, M.H., Suh, S.W.(1995) FEBS Lett 377: 150-154
- PubMed: 8543039 Search on PubMed
- DOI: https://doi.org/10.1016/0014-5793(95)01331-8
- Primary Citation Related Structures:
1KCT - PubMed Abstract:
The crystal structure of a recombinant human alpha 1-antitrypsin, in the uncleaved and uncomplexed state, has been determined by X-ray crystallographic methods and refined to an R-factor of 18.4% for 8.0-3.46 A data with good stereochemistry. This structure provides the first view at the inhibitory loop and the central beta-sheet A of the uncleaved alpha 1-antitrypsin. The reactive loop takes a distorted helical conformation and no pre-insertion of two residues in the reactive loop into the beta-sheet A is observed. The present structure is largely in agreement with the model predicted by Engh, Wright, and Huber [Prot. Eng. 3 (1990) 469-477].
- Department of Chemistry, Seoul National University, South Korea.
Organizational Affiliation: