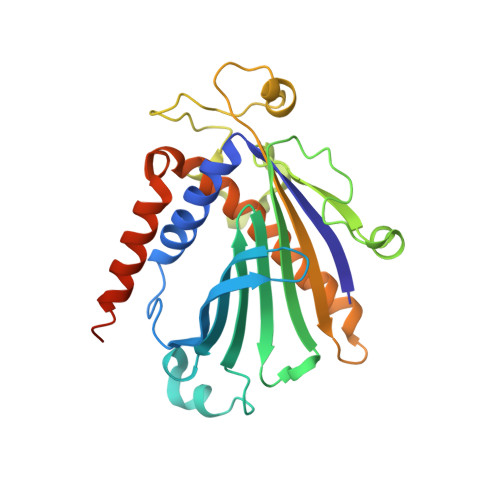
Structure of apo-phosphatidylinositol transfer protein alpha provides insight into membrane association.
Schouten, A., Agianian, B., Westerman, J., Kroon, J., Wirtz, K.W., Gros, P.(2002) EMBO J 21: 2117-2121
- PubMed: 11980708 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/21.9.2117
- Primary Citation Related Structures:
1KCM - PubMed Abstract:
Phosphatidylinositol transfer protein alpha (PITP alpha) is a ubiquitous and highly conserved protein in multicellular eukaryotes that catalyzes the exchange of phospholipids between membranes in vitro and participates in cellular phospholipid metabolism, signal transduction and vesicular trafficking in vivo. Here we report the three-dimensional crystal structure of a phospholipid-free mouse PITP alpha at 2.0 A resolution. The structure reveals an open conformation characterized by a channel running through the protein. The channel is created by opening the phospholipid-binding cavity on one side by displacement of the C-terminal region and a hydrophobic lipid exchange loop, and on the other side by flattening of the central beta-sheet. The relaxed conformation is stabilized at the proposed membrane association site by hydrophobic interactions with a crystallographically related molecule, creating an intimate dimer. The observed open conformer is consistent with a membrane-bound state of PITP and suggests a mechanism for membrane anchoring and the presentation of phosphatidylinositol to kinases and phospholipases after its extraction from the membrane. Coordinates have been deposited in the Protein Data Bank (accession No. 1KCM).
- Department of Crystal and Structural Chemistry, Bijvoet Center for Biomolecular Research, Utrecht University, Padualaan 8,NL-3584 CH Utrecht, The Netherlands.
Organizational Affiliation: