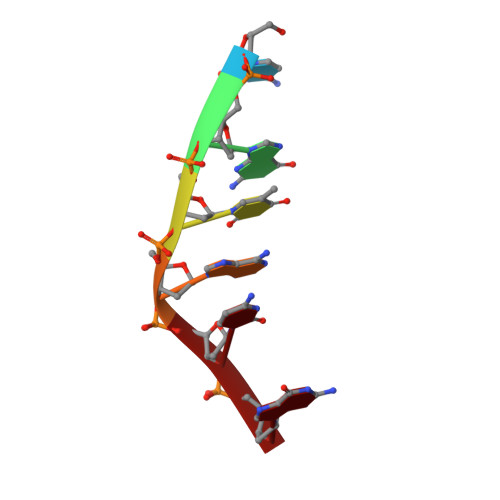
Crystal structure of 9-amino-N-[2-(4-morpholinyl)ethyl]-4-acridinecarboxamide bound to d(CGTACG)2: implications for structure-activity relationships of acridinecarboxamide topoisomerase poisons.
Adams, A., Guss, J.M., Denny, W.A., Wakelin, L.P.(2002) Nucleic Acids Res 30: 719-725
- PubMed: 11809884 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/30.3.719
- Primary Citation Related Structures:
1KCI - PubMed Abstract:
The structure of the complex formed between d(CGTACG)2 and 9-amino-N-[2-(4-morpholinyl)ethyl]-4-acridinecarboxamide, an inactive derivative of the antitumour agents N-[2-(dimethylamino)ethyl]acridine-4-carboxamide (DACA) and 9-amino-DACA, has been solved to a resolution of 1.8 A using X-ray crystallography. The complex crystallises in the space group P6(4 )and the final structure has an overall R factor of 21.9%. A drug molecule intercalates between each of the CpG dinucleotide steps with its side chain lying in the major groove, and its protonated morpholino nitrogen partially occupying positions close to the N7 and O6 atoms of guanine G2. The morpholino group is disordered, the major conformer adopting a twisted boat conformation that makes van der Waals contact with the O4 oxygen of thymine T3. A water molecule forms bridging hydrogen bonds between the 4-carboxamide NH and the phosphate group of guanine G2. Sugar rings are found in alternating C3'-exo/C2'-endo conformations except for cytosine C1 which is C3'-endo. Intercalation perturbs helix winding throughout the hexanucleotide compared with B-DNA, steps 1 and 2 being unwound by 10 and 8 degrees, respectively, while the central TpA step is overwound by 11 degrees. An additional drug molecule lies at the end of each DNA helix linking it to the next duplex to form a continuously stacked structure. The protonated morpholino nitrogen of this 'end-stacked' drug hydrogen bonds to the N7 atom of guanine G6, and its conformationally disordered morpholino ring forms a C-H...O hydrogen bond with the guanine O6 oxygen. In both drug molecules the 4-carboxamide group is internally hydrogen bonded to the protonated N10 atom of the acridine ring. We discuss our findings with respect to the potential role played by the interaction of the drug side chain and the topoisomerase II protein in the poisoning of topoisomerase activity by the acridinecarboxamides.
- Department of Biochemistry, University of Sydney, Sydney, NSW 2006, Australia. a.adams@biochem.usyd.edu.au
Organizational Affiliation: