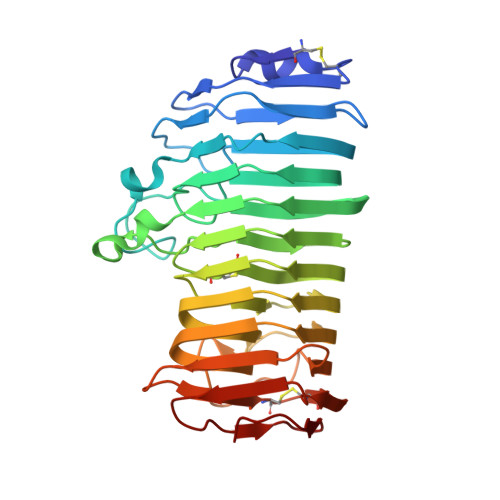
Active-site architecture of endopolygalacturonase I from Stereum purpureum revealed by crystal structures in native and ligand-bound forms at atomic resolution.
Shimizu, T., Nakatsu, T., Miyairi, K., Okuno, T., Kato, H.(2002) Biochemistry 41: 6651-6659
- PubMed: 12022868 Search on PubMed
- DOI: https://doi.org/10.1021/bi025541a
- Primary Citation Related Structures:
1K5C, 1KCC, 1KCD - PubMed Abstract:
Crystal structures of endopolygalacturonase from Stereum purpureum were solved in native and two galacturonic acid complex states at atomic resolution. Endopolygalacturonase catalyzes the hydrolysis of alpha-1,4-glycosidic linkage of polygalacturonate in pectin. The native structure was determined by the multiple wavelength anomalous dispersion method and was refined anisotropically with SHELXL-97, with an R factor of 11.4% and an R(free) factor of 14.0% at 0.96 A resolution. The enzyme folds into a right-handed parallel beta-helix with 10 complete turns. The crystal structures of its binary complex with one D-galacturonate and its ternary complex with two D-galacturonates were also determined to identify the substrate binding site at 1.0 and 1.15 A resolutions, respectively. In the binary complex, one beta-D-galactopyranuronate was found in the +1 subsite, thus proving the strong affinity of the +1 subsite expected from the bond cleavage frequency on oligogalacturonates. In the ternary complex, an additional beta-D-galactofuranuronate was found in the -1 subsite. In both subsites, the recognition of the galacturonate carboxy group is important in galacturonate binding. In the +1 subsite, the carboxy group interacts with three basic residues, His195, Arg226, and Lys228, which were conserved in all endopolygalacturonases. In the -1 subsite, the unique nonprolyl cis-peptide bond is believed to be involved in binding the carboxy group of the substrate. The active site architecture of the complexes provides insight into the mechanism of inverting glycosyl hydrolases and also sheds light on the basis of the differences between the family 28 and the other inverting glycosyl hydrolases.
- Kinetic Crystallography Research Team, Membrane Dynamics Research Group, RIKEN, Harima Institute at SPring-8, 1-1-1 Kouto, Mikazuki-cho, Sayo-gun, Hyogo 679-5148 Japan.
Organizational Affiliation: