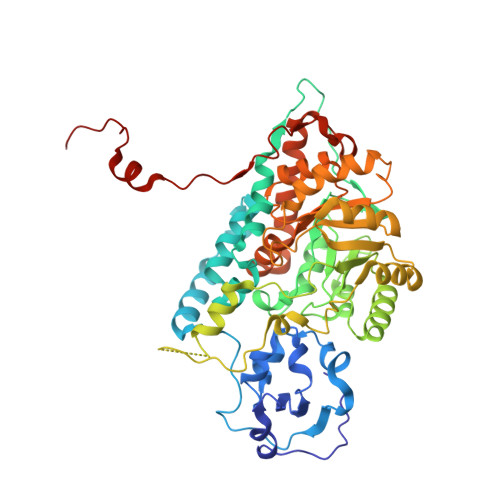
Crystallographic study of the recombinant flavin-binding domain of Baker's yeast flavocytochrome b(2): comparison with the intact wild-type enzyme.
Cunane, L.M., Barton, J.D., Chen, Z.W., Welsh, F.E., Chapman, S.K., Reid, G.A., Mathews, F.S.(2002) Biochemistry 41: 4264-4272
- PubMed: 11914072 Search on PubMed
- Primary Citation Related Structures:
1KBI, 1KBJ - PubMed Abstract:
Flavocytochrome b(2) catalyzes the oxidation of L-lactate to pyruvate and the transfer of electrons to cytochrome c. The enzyme consists of a flavin-binding domain, which includes the active site for lacate oxidation, and a b(2)-cytochrome domain, required for efficient cytochrome c reduction. To better understand the structure and function of intra- and interprotein electron transfer, we have determined the crystal structure of the independently expressed flavin-binding domain of flavocytochrome b(2) to 2.50 A resolution and compared this with the structure of the intact enzyme, redetermined at 2.30 A resolution, both structures being from crystals cooled to 100 K. Whereas there is little overall difference between these structures, we do observe significant local changes near the interface region, some of which impact on amino acid side chains, such as Arg289, that have been shown previously to have an important role in catalysis. The disordered loop region found in flavocytochrome b(2) and its close homologues remain unresolved in frozen crystals of the flavin-binding domain, implying that the presence of the b(2)-cytochrome domain is not responsible for this positional disorder. The flavin-binding domain interacts poorly with cytochrome c, but we have introduced acidic residues in the interdomain interface region with the aim of enhancing cytochrome c binding. While the mutations L199E and K201E within the flavin-binding domain resulted in unimpaired lactate dehydrogenase activity, they failed to enhance electron-transfer rates with cytochrome c. This is most likely due to the disordered loop region obscuring all or part of the surface having the potential for productive interaction with cytochrome c.
- Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, Missouri 63110, USA.
Organizational Affiliation: