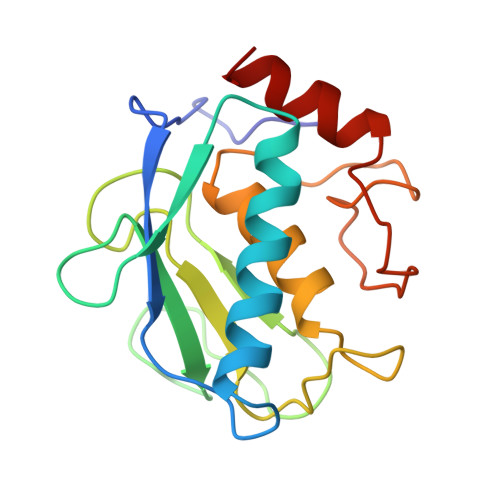
1.8-A crystal structure of the catalytic domain of human neutrophil collagenase (matrix metalloproteinase-8) complexed with a peptidomimetic hydroxamate primed-side inhibitor with a distinct selectivity profile.
Betz, M., Huxley, P., Davies, S.J., Mushtaq, Y., Pieper, M., Tschesche, H., Bode, W., Gomis-Ruth, F.X.(1997) Eur J Biochem 247: 356-363
- PubMed: 9249047 Search on PubMed
- DOI: https://doi.org/10.1111/j.1432-1033.1997.00356.x
- Primary Citation Related Structures:
1KBC - PubMed Abstract:
Matrix metalloproteinases (MMP) are zinc endopeptidases involved in tissue remodelling. They have been implicated in a series of pathologies, including cancer, arthritis, joint destruction and Alzheimer's disease. Human neutrophil collagenase represents one of the three interstitial collagenases that cleave triple-helical collagen of type I, II and III. Its catalytic domain (residues Phe79-Gly242) has been heterologously expressed in Escherichia coli and crystallized as a non-covalent complex with the hydroxamate inhibitor BB-1909, which has distinct selectivity against different MMP, in a crystal form. The crystal structure, refined to 0.18-nm resolution, shows that BB-1909 is a right-hand-side inhibitor that binds to the S1'-S3' subsites and coordinates to the catalytic Zn2+ in a bidentate manner via the hydroxyl and carbonyl oxygen atoms of the hydroxamate group in a similar manner to batimastat. The collagenase/BB-1909 complex is described in detail and compared with the collagenase/batimastat complex. These studies provide information on MMP specificity and thus may assist the development of more-selective MMP inhibitors.
- Max-Planck-Institut für Biochemie, Abteilung Strukturforschung, Planegg-Martinsried, Germany.
Organizational Affiliation: