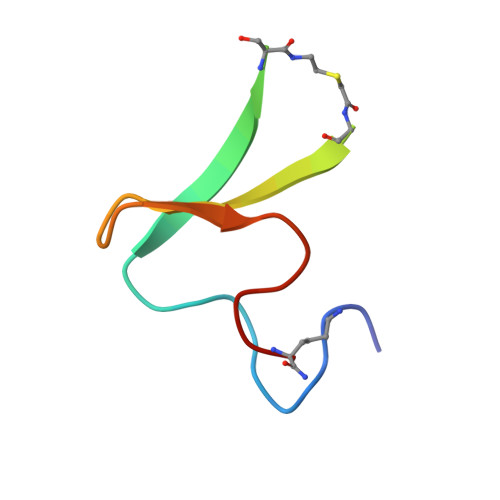
Using flexible loop mimetics to extend phi-value analysis to secondary structure interactions.
Ferguson, N., Pires, J.R., Toepert, F., Johnson, C.M., Pan, Y.P., Volkmer-Engert, R., Schneider-Mergener, J., Daggett, V., Oschkinat, H., Fersht, A.(2001) Proc Natl Acad Sci U S A 98: 13008-13013
- PubMed: 11687614 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.221467398
- Primary Citation Related Structures:
1K5R - PubMed Abstract:
Chemical synthesis allows the incorporation of nonnatural amino acids into proteins that may provide previously untried probes of their folding pathway and thermodynamic stability. We have used a flexible thioether linker as a loop mimetic in the human yes kinase-associated protein (YAP 65) WW domain, a three-stranded, 44-residue, beta-sheet protein. This linkage avoids problems of incorporating sequences that constrain loops to the extent that they significantly change the nature of the denatured state with concomitant effects on the folding kinetics. An NMR solution structure shows that the thioether linker had little effect on the global fold of the domain, although the loop is apparently more dynamic. The thioether variants are destabilized by up to 1.4 kcal/mol (1 cal = 4.18 J). Preliminary Phi-value analysis showed that the first loop is highly structured in the folding transition state, and the second loop is essentially unstructured. These data are consistent with results from simulated unfolding and detailed protein-engineering studies of structurally homologous WW domains. Previously, Phi-value analysis was limited to studying side-chain interactions. The linkers used here extend the protein engineering method directly to secondary-structure interactions.
- Medical Research Council, Centre for Protein Engineering, Hills Road, Cambridge CB2 2QH, United Kingdom.
Organizational Affiliation: