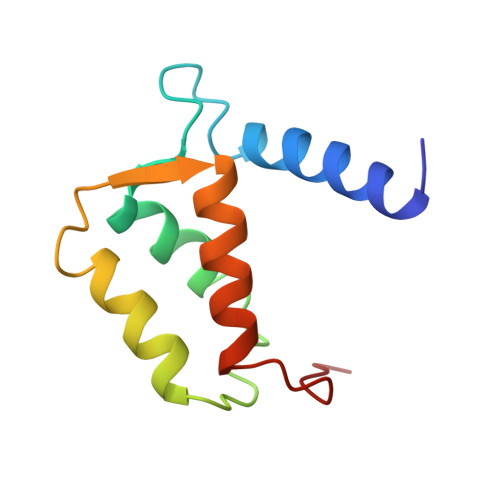
Three-dimensional solution structure of the calcium-signaling protein apo-S100A1 as determined by NMR.
Rustandi, R.R., Baldisseri, D.M., Inman, K.G., Nizner, P., Hamilton, S.M., Landar, A., Landar, A., Zimmer, D.B., Weber, D.J.(2002) Biochemistry 41: 788-796
- PubMed: 11790100 Search on PubMed
- DOI: https://doi.org/10.1021/bi0118308
- Primary Citation Related Structures:
1K2H - PubMed Abstract:
S100A1, a member of the S100 protein family, is an EF-hand containing Ca(2+)-binding protein (93 residues per subunit) with noncovalent interactions at its dimer interface. Each subunit of S100A1 has four alpha-helices and a small antiparallel beta-sheet consistent with two helix-loop-helix calcium-binding domains [Baldiserri et al. (1999) J. Biomol. NMR 14, 87-88]. In this study, the three-dimensional structure of reduced apo-S100A1 was determined by NMR spectroscopy using a total of 2220 NOE distance constraints, 258 dihedral angle constraints, and 168 backbone hydrogen bond constraints derived from a series of 2D, 3D, and 4D NMR experiments. The final structure was found to be globular and compact with the four helices in each subunit aligning to form a unicornate-type four-helix bundle. Intermolecular NOE correlations were observed between residues in helices 1 and 4 from one subunit to residues in helices 1' and 4' of the other subunit, respectively, consistent with the antiparallel alignment of the two subunits to form a symmetric X-type four-helix bundle as found for other members of the S100 protein family. Because of the similarity of the S100A1 dimer interface to that found for S100B, it was possible to calculate a model of the S100A1/B heterodimer. This model is consistent with a number of NMR chemical shift changes observed when S100A1 is titrated into a sample of (15)N-labeled S100B. Helix 3 (and 3') of S100A1 was found to have an interhelical angle of -150 degrees with helix 4 (and 4') in the apo state. This crossing angle is quite different (>50 degrees ) from that typically found in other EF-hand containing proteins such as apocalmodulin and apotroponin C but more similar to apo-S100B, which has an interhelical angle of -166 degrees. As with S100B, it is likely that the second EF-hand of apo-S100A1 reorients dramatically upon the addition of Ca(2+), which can explain the Ca(2+) dependence that S100A1 has for binding several of its biological targets.
- Department of Biochemistry and Molecular Biology, University of Maryland School of Medicine, 108 North Greene Street, Baltimore, Maryland 21201, USA.
Organizational Affiliation: