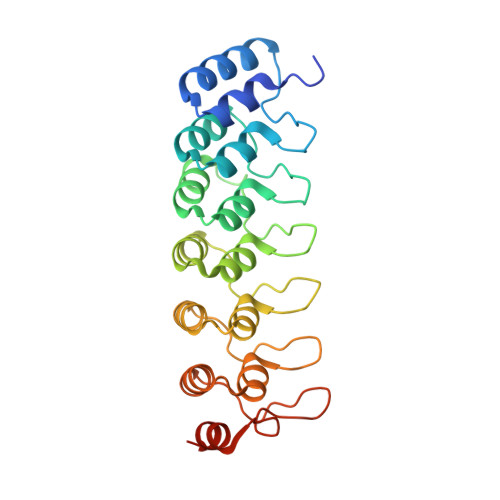
Crystal structure of the ankyrin repeat domain of Bcl-3: a unique member of the IkappaB protein family.
Michel, F., Soler-Lopez, M., Petosa, C., Cramer, P., Siebenlist, U., Muller, C.W.(2001) EMBO J 20: 6180-6190
- PubMed: 11707390 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/20.22.6180
- Primary Citation Related Structures:
1K1A, 1K1B - PubMed Abstract:
IkappaB proteins associate with the transcription factor NF-kappaB via their ankyrin repeat domain. Bcl-3 is an unusual IkappaB protein because it is primarily nucleoplasmic and can lead to enhanced NF-kappaB-dependent transcription, unlike the prototypical IkappaB protein IkappaBalpha, which inhibits NF-kappaB activity by retaining it in the cytoplasm. Here we report the 1.9 A crystal structure of the ankyrin repeat domain of human Bcl-3 and compare it with that of IkappaBalpha bound to NF-kappaB. The two structures are highly similar over the central ankyrin repeats but differ in the N-terminal repeat and at the C-terminus, where Bcl-3 contains a seventh repeat in place of the acidic PEST region of IkappaBalpha. Differences between the two structures suggest why Bcl-3 differs from IkappaBalpha in selectivity towards various NF-kappaB species, why Bcl-3 but not IkappaBalpha can associate with its NF-kappaB partner bound to DNA, and why two molecules of Bcl-3 but only one of IkappaBalpha can bind to its NF-kappaB partner. Comparison of the two structures thus provides an insight into the functional diversity of IkappaB proteins.
- European Molecular Biology Laboratory, Grenoble Outstation BP 181, 38042 Grenoble, Cedex 9, France.
Organizational Affiliation: