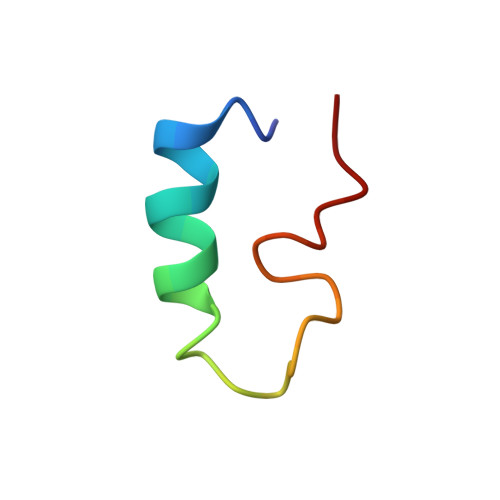
Nuclear magnetic resonance structures of the zinc finger domain of human DNA polymerase-alpha.
Evanics, F., Maurmann, L., Yang, W.W., Bose, R.N.(2003) Biochim Biophys Acta 1651: 163-171
- PubMed: 14499601 Search on PubMed
- DOI: https://doi.org/10.1016/s1570-9639(03)00266-8
- Primary Citation Related Structures:
1K0P, 1K18, 1N5G - PubMed Abstract:
The carboxy terminus of the human DNA polymerase-alpha contains a zinc finger motif. Three-dimensional structures of this motif containing 38 amino acid residues, W L I C E E P T C R N R T R H L P L Q F S R T G P L C P A C M K A T L Q P E, were determined by nuclear magnetic resonance (NMR) spectroscopy. The structures reveal an alpha-helix-like domain at the amino terminus, extending 13 residues from L2 through H15 with an interruption at the sixth residue. The helix region is followed by three turns (H15-L18, T23-L26 and L26-A29), all of which involve proline. The first turn appears to be type III, judging by the dihedral angles. The second and third turns appear to be atypical. A second, shorter helix is formed at the carboxy terminus extending from C30 through L35. A fourth type III turn starting at L35 was also observed in the structure. Proline serves as the third residue of all the turns. Four cysteine residues, two located at the beginning of the helix at the N-terminus and two at the carboxy end, are coordinated to Zn(II), facilitating the formation of a loop. One of the cysteines at the carboxy terminus is part of the atypical turn, while the other is the part of the short helix. These structural features are consistent with the circular dichroism (CD) measurements which indicate the presence of 45% helix, 11% beta turns and 19% non-ordered secondary structures. The zinc finger motif described here is different from those observed for C(4), C(2)H(2), and C(2)HC modules reported in the literature. In particular, polymerase-alpha structures exhibit helix-turn-helix motif while most zinc finger proteins show anti-parallel sheet and helix. Several residues capable of binding DNA, T, R, N, and H are located in the helical region. These structural features imply that the zinc finger motif is most likely involved in binding DNA prior to replication, presumably through the helical region. These results are discussed in the context of other eukaryotic and prokaryotic DNA polymerases belonging to the polymerase B family.
- Department of Chemistry and Biochemistry, Northern Illinois University, DeKalb, IL 60115, USA.
Organizational Affiliation: