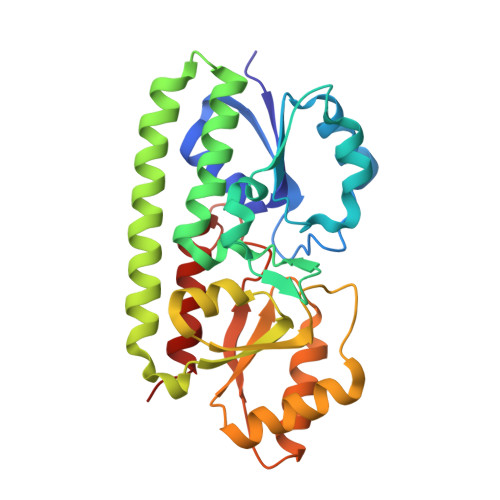
The crystal structure of Zn(II)-free Treponema pallidum TroA, a periplasmic metal-binding protein, reveals a closed conformation.
Lee, Y.H., Dorwart, M.R., Hazlett, K.R., Deka, R.K., Norgard, M.V., Radolf, J.D., Hasemann, C.A.(2002) J Bacteriol 184: 2300-2304
- PubMed: 11914363 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.184.8.2300-2304.2002
- Primary Citation Related Structures:
1K0F - PubMed Abstract:
We previously demonstrated that Treponema pallidum TroA is a periplasmic metal-binding protein (MBP) with a distinctive alpha-helical backbone. To better understand the mechanisms of metal binding and release by TroA, we determined the crystal structure of the apoprotein at a resolution of 2.5 A and compared it to that of the Zn(II)-bound form (Protein Data Bank accession code 1toa). apo-TroA shows a conformation even more closed than that of its Zn(II)-bound counterpart due to a 4 degrees tilt of the C-terminal domain (residues 190 through 308) about an axis parallel to the poorly flexible backbone helix. This domain tilting pushes two loops (residues 248 through 253 and 277 through 286) towards the metal-binding site by more than 1 A, resulting in an unfavorable interaction of I251 with D66. To avoid this contact, D66 shifts towards H68, one of the four Zn(II)-coordinating residues. The approach of this negative charge coincides with the flipping of the imidazole side chain of H68, resulting in the formation of a new hydrogen bond. The conformational change of H68, along with a slight rearrangement of D279, a C-terminal domain Zn(II)-coordinating residue, distorts the metal-binding site geometry, presumably causing the release of the bound metal ion. Ligand binding and release by TroA, and presumably by other members of the MBP cluster, differs from the "Venus flytrap" mechanism utilized by bacterial nonmetal solute-binding receptors.
- Department of Biochemistry, The University of Missouri-Columbia, Columbia, Missouri 65211, USA.
Organizational Affiliation: