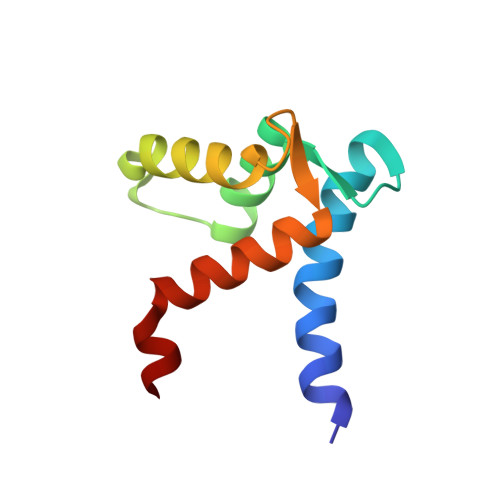
A structural basis for S100 protein specificity derived from comparative analysis of apo and Ca(2+)-calcyclin
Maler, L., Sastry, M., Chazin, W.J.(2002) J Mol Biology 317: 279-290
- PubMed: 11902843 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2002.5421
- Primary Citation Related Structures:
1JWD - PubMed Abstract:
Calcyclin is a homodimeric protein belonging to the S100 subfamily of EF-hand Ca(2+)-binding proteins, which function in Ca(2+) signal transduction processes. A refined high-resolution solution structure of Ca(2+)-bound rabbit calcyclin has been determined by heteronuclear solution NMR. In order to understand the Ca(2+)-induced structural changes in S100 proteins, in-depth comparative structural analyses were used to compare the apo and Ca(2+)-bound states of calcyclin, the closely related S100B, and the prototypical Ca(2+)-sensor protein calmodulin. Upon Ca(2+) binding, the position and orientation of helix III in the second EF-hand is altered, whereas the rest of the protein, including the dimer interface, remains virtually unchanged. This Ca(2+)-induced structural change is much less drastic than the "opening" of the globular EF-hand domains that occurs in classical Ca(2+) sensors, such as calmodulin. Using homology models of calcyclin based on S100B, a binding site in calcyclin has been proposed for the N-terminal domain of annexin XI and the C-terminal domain of the neuronal calcyclin-binding protein. The structural basis for the specificity of S100 proteins is discussed in terms of the variation in sequence of critical contact residues in the common S100 target-binding site.
- Department of Biochemistry and Biophysics, Arrhenius Laboratory, Stockholm University, Sweden.
Organizational Affiliation: