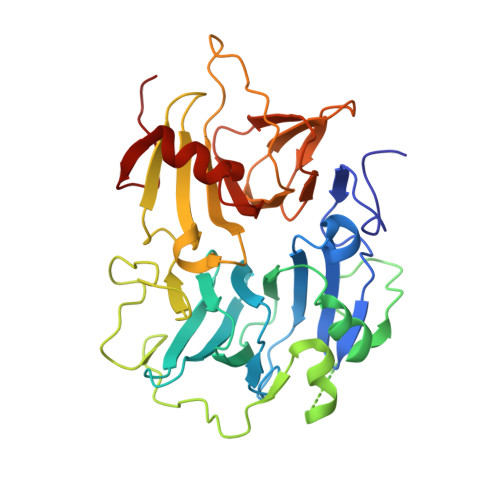
Crystal structure of the copper-containing quercetin 2,3-dioxygenase from Aspergillus japonicus.
Fusetti, F., Schroter, K.H., Steiner, R.A., van Noort, P.I., Pijning, T., Rozeboom, H.J., Kalk, K.H., Egmond, M.R., Dijkstra, B.W.(2002) Structure 10: 259-268
- PubMed: 11839311 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00704-9
- Primary Citation Related Structures:
1JUH - PubMed Abstract:
Quercetin 2,3-dioxygenase is a copper-containing enzyme that catalyzes the insertion of molecular oxygen into polyphenolic flavonols. Dioxygenation catalyzed by iron-containing enzymes has been studied extensively, but dioxygenases employing other metal cofactors are poorly understood. We determined the crystal structure of quercetin 2,3-dioxygenase at 1.6 A resolution. The enzyme forms homodimers, which are stabilized by an N-linked heptasaccharide at the dimer interface. The mononuclear type 2 copper center displays two distinct geometries: a distorted tetrahedral coordination, formed by His66, His68, His112, and a water molecule, and a distorted trigonal bipyramidal environment, which additionally comprises Glu73. Manual docking of the substrate quercetin into the active site showed that the different geometries of the copper site might be of catalytic importance.
- Laboratory of Biophysical Chemistry, Department of Chemistry, University of Groningen, Nijenborgh 4, 9747 AG Groningen, The Netherlands.
Organizational Affiliation: