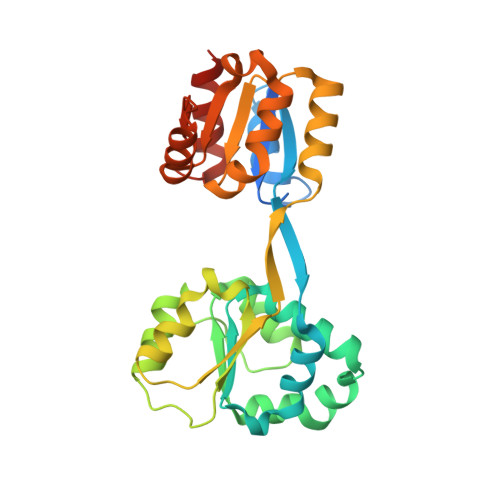
Crystal structure of human uroporphyrinogen III synthase.
Mathews, M.A., Schubert, H.L., Whitby, F.G., Alexander, K.J., Schadick, K., Bergonia, H.A., Phillips, J.D., Hill, C.P.(2001) EMBO J 20: 5832-5839
- PubMed: 11689424 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/20.21.5832
- Primary Citation Related Structures:
1JR2 - PubMed Abstract:
Uroporphyrinogen III synthase, U3S, the fourth enzyme in the porphyrin biosynthetic pathway, catalyzes cyclization of the linear tetrapyrrole, hydroxymethylbilane, to the macrocyclic uroporphyrino gen III, which is used in several different pathways to form heme, siroheme, chlorophyll, F(430) and vitamin B(12). U3S activity is essential in all organisms, and decreased activity in humans leads to the autosomal recessive disorder congenital erythropoetic porphyria. We have determined the crystal structure of recombinant human U3S at 1.85 A resolution. The protein folds into two alpha/beta domains connected by a beta-ladder. The active site appears to be located between the domains, and variations in relative domain positions observed between crystallographically independent molecules indicates the presence of flexibility that may be important in the catalytic cycle. Possible mechanisms of catalysis were probed by mutating each of the four invariant residues in the protein that have titratable side chains. Additionally, six other highly conserved and titratable side chains were also mutated. In no case, however, did one of these mutations abolish enzyme activity, suggesting that the mechanism does not require acid/base catalysis.
- Department of Biochemistry, University of Utah School of Medicine, 50 N. Medical Drive, Salt Lake City, UT 84132, USA.
Organizational Affiliation: