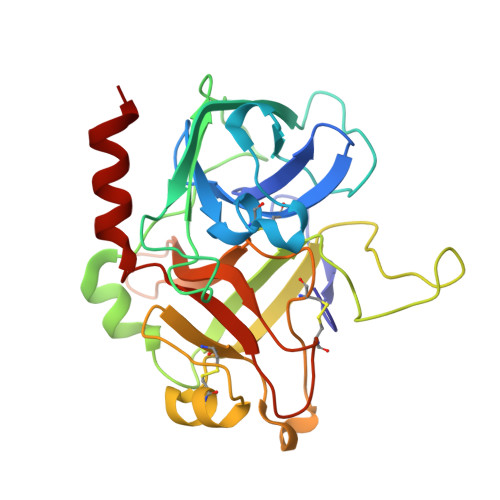
The molecular basis of thrombin allostery revealed by a 1.8A structure of the slow form
Huntington, J.A., Esmon, C.T.(2003) Structure 11: 469-479
- PubMed: 12679024 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(03)00049-2
- Primary Citation Related Structures:
1JOU - PubMed Abstract:
Thrombin participates in its own positive and negative feedback loops, and its allosteric state helps determine the hemostatic balance. Here we present the 1.8 A crystallographic structure of S195A thrombin in two conformational states: active site occupied and active site free. The active site-occupied form shows how thrombin can accommodate substrates, such as protein C. The active site-free form is in a previously unobserved closed conformation of thrombin, which satisfies all the conditions of the so-called "slow" form. A mechanism of allostery is revealed, which relies on the concerted movement of the disulphide bond between Cys168 and 182 and aromatic residues Phe227, Trp215, and Trp60d. These residues constitute an allosteric switch, which is flipped directly through sodium binding, resulting in the fast form with an open active site.
- Department of Haematology, University of Cambridge, Cambridge Institute for Medical Research, Wellcome Trust/MRC Building, Hills Road, CB2 2XY, Cambridge, United Kingdom. jah52@cam.ac.uk
Organizational Affiliation: