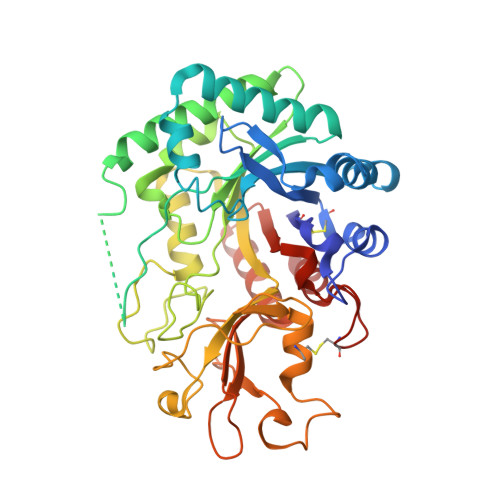
Crystal structure of imaginal disc growth factor-2. A member of a new family of growth-promoting glycoproteins from Drosophila melanogaster.
Varela, P.F., Llera, A.S., Mariuzza, R.A., Tormo, J.(2002) J Biological Chem 277: 13229-13236
- PubMed: 11821393 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M110502200
- Primary Citation Related Structures:
1JND, 1JNE - PubMed Abstract:
Imaginal disc growth factor-2 (IDGF-2) is a member of a recently described family of Drosophila melanogaster-soluble polypeptide growth factors that promote cell proliferation in imaginal discs. Although their precise mode of action has not been established, IDGFs cooperate with insulin in stimulating the growth of imaginal disc cells. We report the crystal structure of IDGF-2 at 1.3-A resolution. The structure shows the classical (betaalpha)(8) barrel-fold of family 18 glycosyl hydrolases, with an insertion of an alpha + beta domain similar to that of Serratia marcescens chitinases A and B. However, amino acid substitutions in the consensus catalytic sequence of chitinases give IDGF-2 a less negatively charged environment in its putative ligand-binding site and preclude the nucleophilic attack mechanism of chitin hydrolysis. Particularly important is the replacement of Glu by Gln at position 132, which has been shown to abolish enzymatic activity in chitinases. Nevertheless, a modest conservation of residues that participate in oligosaccharide recognition suggests that IDGF-2 could bind carbohydrates, assuming several conformational changes to open the partially occluded binding site. Thus, IDGFs may have evolved from chitinases to acquire new functions as growth factors, interacting with cell surface glycoproteins implicated in growth-promoting processes, such as the Drosophila insulin receptor.
- Center for Advanced Research in Biotechnology, W. M. Keck Laboratory of Structural Biology, University of Maryland Biotechnology Institute, Rockville, MD 20850, USA.
Organizational Affiliation: