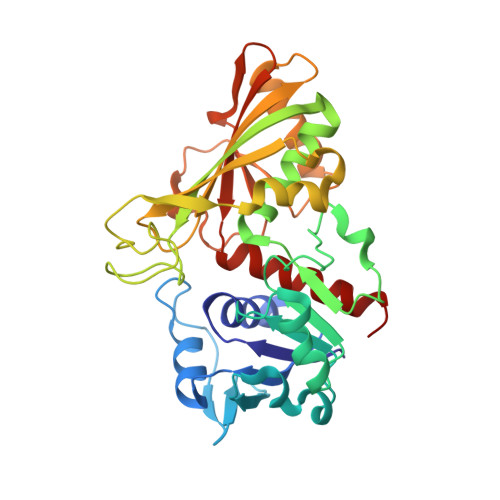
Crystal structure of the non-regulatory A(4 )isoform of spinach chloroplast glyceraldehyde-3-phosphate dehydrogenase complexed with NADP.
Fermani, S., Ripamonti, A., Sabatino, P., Zanotti, G., Scagliarini, S., Sparla, F., Trost, P., Pupillo, P.(2001) J Mol Biology 314: 527-542
- PubMed: 11846565 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.5172
- Primary Citation Related Structures:
1JN0 - PubMed Abstract:
Here, we report the first crystal structure of a photosynthetic glyceraldehyde-3-phosphate dehydrogenase (GAPDH) complexed with NADP. The enzyme, purified from spinach chloroplasts, is constituted of a single type of subunit (A) arranged in homotetramers. It shows non-regulated NADP-dependent and NAD-dependent activities, with a preference for NADP. The structure has been solved to 3.0 A resolution by molecular replacement. The crystals belong to space group C222 with three monomers in the asymmetric unit. One of the three monomers generates a tetramer using the space group 222 point symmetry and a very similar tetramer is generated by the other two monomers, related by a non-crystallographic symmetry, using a crystallographic 2-fold axis. The protein reveals a large structural homology with known GAPDHs both in the cofactor-binding domain and in regions of the catalytic domain. Like all other GAPDHs investigated so far, the A(4)-GAPDH belongs to the Rossmann fold family of dehydrogenases. However, unlike most dehydrogenases of this family, the adenosine 2'-phosphate group of NADP does not form a salt-bridge with any positively charged residue in its surroundings, being instead set in place by hydrogen bonds with a threonine residue belonging to the Rossmann fold and a serine residue located in the S-loop of a symmetry-related monomer. While increasing our knowledge of an important photosynthetic enzyme, these results contribute to a general understanding of NADP versus NAD recognition in pyridine nucleotide-dependent enzymes. Although the overall structure of A(4)-GAPDH is similar to that of the cytosolic GAPDH from bacteria and eukaryotes, the chloroplast tetramer is peculiar, in that it can actually be considered a dimer of dimers, since monomers are bound in pairs by a disulphide bridge formed across Cys200 residues. This bridge is not found in other cytosolic or chloroplast GAPDHs from animals, bacteria, or plants other than spinach.
- Dipartimento di Chimica "G. Ciamician", Università di Bologna, via Selmi 2, Bologna, 40126, Italia. fermani@ciam.unibo.it
Organizational Affiliation: