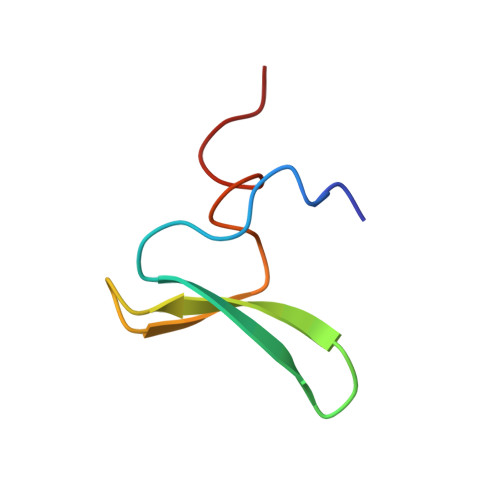
Solution structures of the YAP65 WW domain and the variant L30 K in complex with the peptides GTPPPPYTVG, N-(n-octyl)-GPPPY and PLPPY and the application of peptide libraries reveal a minimal binding epitope.
Pires, J.R., Taha-Nejad, F., Toepert, F., Ast, T., Hoffmuller, U., Schneider-Mergener, J., Kuhne, R., Macias, M.J., Oschkinat, H.(2001) J Mol Biology 314: 1147-1156
- PubMed: 11743730 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.5199
- Primary Citation Related Structures:
1JMQ, 1K9Q, 1K9R - PubMed Abstract:
The single mutation L30 K in the Hu-Yap65 WW domain increased the stability of the complex with the peptide GTPPPPYTVG (K(d)=40(+/-5) microM). Here we report the refined solution structure of this complex by NMR spectroscopy and further derived structure-activity relationships by using ligand peptide libraries with truncated sequences and a substitution analysis that yielded acetyl-PPPPY as the smallest high-affinity binding peptide (K(d)=60 microM). The structures of two new complexes with weaker binding ligands chosen based on these results (N-(n-octyl)-GPPPYNH(2) and Ac-PLPPY) comprising the wild-type WW domain of Hu-Yap65 were determined. Comparison of the structures of the three complexes were useful for identifying the molecular basis of high-affinity: hydrophobic and specific interactions between the side-chains of Y28 and W39 and P5' and P4', respectively, and hydrogen bonds between T37 (donnor) and P5' (acceptor) and between W39 (donnor) and T2' (acceptor) stabilize the complex.The structure of the complex L30 K Hu-Yap65 WW domain/GTPPPPYTVG is compared to the published crystal structure of the dystrophin WW domain bound to a segment of the beta-dystroglycan protein and to the solution structure of the first Nedd4 WW domain and its prolin-rich ligand, suggesting that WW sequences bind proline-rich peptides in an evolutionary conserved fashion. The position equivalent to T22 in the Hu-Yap65 WW domain sequence is seen as responsible for differentiation in the binding mode among the WW domains of group I.
- Forschungsinstitut für Molekulare Pharmakologie, Robert-Rössle-Str. 10, 13125 Berlin, Germany. oschkinat@fmp-berlin.de
Organizational Affiliation: