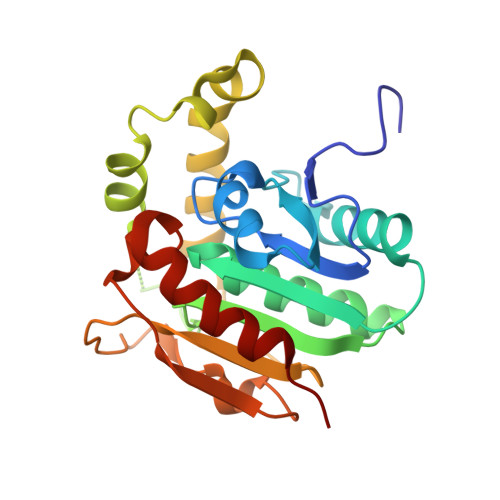
Structural basis for the cyclization of the lipopeptide antibiotic surfactin by the thioesterase domain SrfTE.
Bruner, S.D., Weber, T., Kohli, R.M., Schwarzer, D., Marahiel, M.A., Walsh, C.T., Stubbs, M.T.(2002) Structure 10: 301-310
- PubMed: 12005429 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00716-5
- Primary Citation Related Structures:
1JMK - PubMed Abstract:
Many biologically active natural peptides are synthesized by nonribosomal peptide synthetases (NRPS). Product release is accomplished by dedicated thioesterase (TE) domains, some of which catalyze an intramolecular cyclization to form macrolactone or macrolactam cyclic peptides. The excised 28 kDa SrfTE domain, a member of the alpha/beta hydrolase enzyme family, exhibits a distinctive bowl-shaped hydrophobic cavity that hosts the acylpeptide substrate and tolerates its folding to form a cyclic structure. A substrate analog confirms the substrate binding site and suggests a mechanism for substrate acylation/deacylation. Docking of the peptidyl carrier protein domain immediately preceding SrfTE positions the 4'-phosphopantheinyl prosthetic group that transfers the nascent acyl-peptide chain to SrfTE. The structure provides a basis for understanding the mechanism of acyl-PCP substrate recognition and for the cyclization reaction that results in release of the macrolactone cyclic heptapeptide.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts 02115, USA.
Organizational Affiliation: