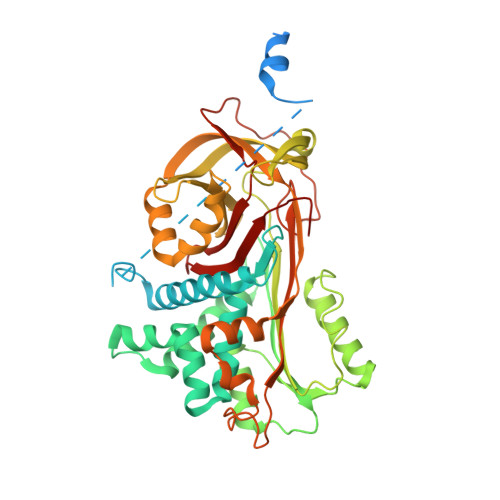
Crystal structures of native and thrombin-complexed heparin cofactor II reveal a multistep allosteric mechanism.
Baglin, T.P., Carrell, R.W., Church, F.C., Esmon, C.T., Huntington, J.A.(2002) Proc Natl Acad Sci U S A 99: 11079-11084
- PubMed: 12169660 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.162232399
- Primary Citation Related Structures:
1JMJ, 1JMO - PubMed Abstract:
The serine proteases sequentially activated to form a fibrin clot are inhibited primarily by members of the serpin family, which use a unique beta-sheet expansion mechanism to trap and destroy their targets. Since the discovery that serpins were a family of serine protease inhibitors there has been controversy as to the role of conformational change in their mechanism. It now is clear that protease inhibition depends entirely on rapid serpin beta-sheet expansion after proteolytic attack. The regulatory advantage afforded by the conformational mobility of serpins is demonstrated here by the structures of native and S195A thrombin-complexed heparin cofactor II (HCII). HCII inhibits thrombin, the final protease of the coagulation cascade, in a glycosaminoglycan-dependent manner that involves the release of a sequestered hirudin-like N-terminal tail for interaction with thrombin. The native structure of HCII resembles that of native antithrombin and suggests an alternative mechanism of allosteric activation, whereas the structure of the S195A thrombin-HCII complex defines the molecular basis of allostery. Together, these structures reveal a multistep allosteric mechanism that relies on sequential contraction and expansion of the central beta-sheet of HCII.
- Department of Haematology, Cambridge Institute for Medical Research, Wellcome Trust/MRC Building, Hills Road, Cambridge CB2 2XY, United Kingdom.
Organizational Affiliation: