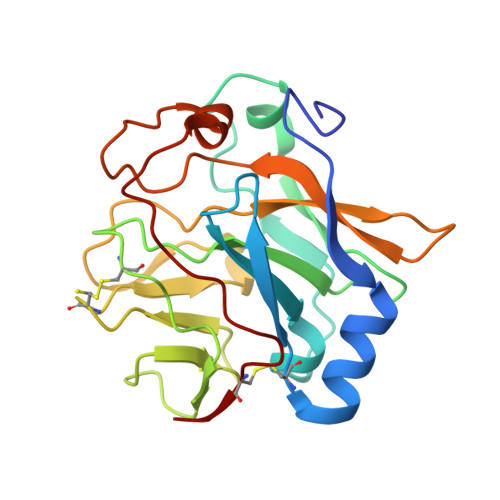
The structure of the soluble domain of an archaeal Rieske iron-sulfur protein at 1.1 A resolution.
Bonisch, H., Schmidt, C.L., Schafer, G., Ladenstein, R.(2002) J Mol Biology 319: 791-805
- PubMed: 12054871 Search on PubMed
- DOI: https://doi.org/10.1016/S0022-2836(02)00323-6
- Primary Citation Related Structures:
1JM1 - PubMed Abstract:
The first crystal structure of an archaeal Rieske iron-sulfur protein, the soluble domain of Rieske iron-sulfur protein II (soxF) from the hyperthermo-acidophile Sulfolobus acidocaldarius, has been solved by multiple wavelength anomalous dispersion (MAD) and has been refined to 1.1 A resolution. SoxF is a subunit of the terminal oxidase supercomplex SoxM in the plasma membrane of S. acidocaldarius that combines features of a cytochrome bc(1) complex and a cytochrome c oxidase. The [2Fe-2S] cluster of soxF is most likely the primary electron acceptor during the oxidation of caldariella quinone by the cytochrome a(587)/Rieske subcomplex. The geometry of the [2Fe-2S] cluster and the structure of the cluster-binding site are almost identical in soxF and the Rieske proteins from eucaryal cytochrome bc(1) and b(6)f complexes, suggesting a strict conservation of the catalytic mechanism. The main domain of soxF and part of the cluster-binding domain, though structurally related, show a significantly divergent structure with respect to topology, non-covalent interactions and surface charges. The divergent structure of soxF reflects a different topology of the soxM complex compared to eucaryal bc complexes and the adaptation of the protein to the extreme ambient conditions on the outer membrane surface of a hyperthermo-acidophilic organism.
- Department of Biosciences at NOVUM, Center for Structural Biochemistry, Karolinska Institutet, Hälsovägen 7-9, S-14157 Huddinge, Sweden.
Organizational Affiliation: