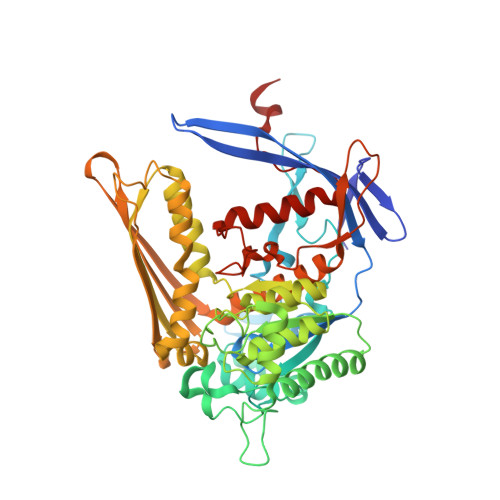
The crystal structure and mechanism of 1-L-myo-inositol- 1-phosphate synthase
Stein, A.J., Geiger, J.H.(2002) J Biological Chem 277: 9484-9491
- PubMed: 11779862 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M109371200
- Primary Citation Related Structures:
1JKF, 1JKI - PubMed Abstract:
1-l-myo-Inositol-1-phosphate synthase catalyzes the conversion of d-glucose 6-phosphate to 1-l-myo-inositol-1-phosphate (MIP), the first and rate-limiting step in the biosynthesis of all inositol-containing compounds. It involves an oxidation, intramolecular aldol cyclization, and reduction. We have determined the first crystal structure of MIP synthase. We present structures of both the NAD-bound enzyme and the enzyme bound to an inhibitor, 2-deoxy-glucitol-6-phosphate. While 58 amino acids are disordered in the unbound form of the enzyme in the vicinity of the active site, the inhibitor nucleates the folding of this domain in a striking example of induced fit, serving to completely encapsulate it within the enzyme. Three helices and a long beta-strand are formed in this process. We postulate a mechanism for the conversion based on the structure of the inhibitor-bound complex.
- Department of Chemistry, Michigan State University, East Lansing, Michigan 48824, USA.
Organizational Affiliation: