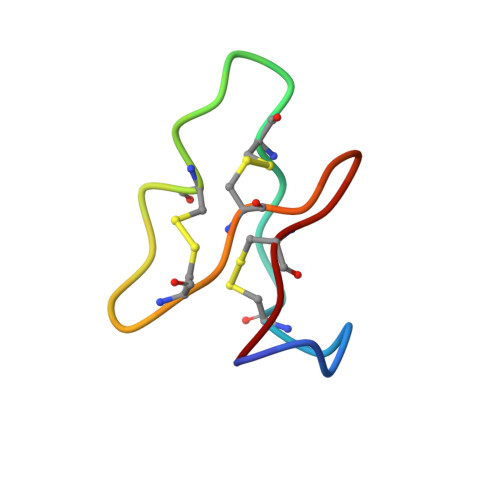
Refined structure and metal binding site of the kalata B1 peptide.
Skjeldal, L., Gran, L., Sletten, K., Volkman, B.F.(2002) Arch Biochem Biophys 399: 142-148
- PubMed: 11888199 Search on PubMed
- DOI: https://doi.org/10.1006/abbi.2002.2769
- Primary Citation Related Structures:
1JJZ, 1K48 - PubMed Abstract:
The cyclic polypeptide kalata B1 from the African plant Oldenlandia affinis DC consists of 29 amino acid residues with three disulfide linkages. In this study we used two-dimensional NMR spectroscopy to investigate the three-dimensional structure of the peptide and to determine the disulfide connectivities. Nuclear Overhauser effects (NOEs) between neighboring beta-protons of the cysteines detected at 750 MHz provided evidence for the disulfide connectivity pattern 5-13, 17-29, and 22-27. These disulfide linkages were confirmed by three-dimensional structures calculated from input constraints derived solely from NOEs without explicit disulfide connectivities. Kalata B1 is insoluble in aqueous solution above pH 3.5, but in a 50-50 water-methanol mixture, it was possible to use natural abundance two-dimensional (15)N-(1)H heteronuclear single quantum coherence spectroscopy to study the hydrophobic peptide from pH 2 to 10. The addition of methanol resulted in no significant structural changes. Although the peptide contains three prolyl residues, no evidence of multiple conformers was detected at any pH. The addition of Mn(2+) to kalata B1 resulted in selective broadening of resonances from Asn 23, Thr 24, and Glu 15; these results suggest that these three residues are involved in a specific metal binding site.
- Department of Chemistry and Biotechnology, Agricultural University of Norway, N-1432 As, Norway.
Organizational Affiliation: