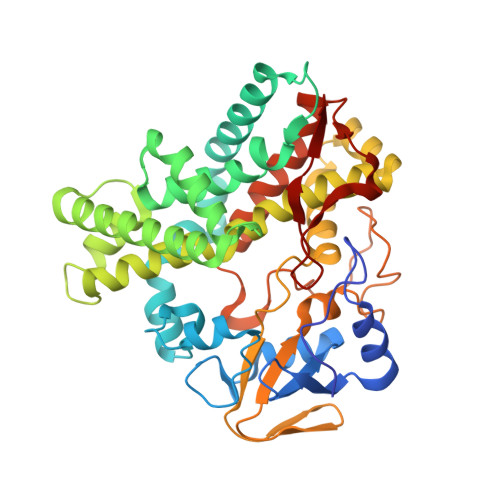
X-ray structure of nitric oxide reductase (cytochrome P450nor) at atomic resolution.
Shimizu, H., Park, S.Y., Shiro, Y., Adachi, S.(2002) Acta Crystallogr D Biol Crystallogr 58: 81-89
- PubMed: 11752781 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444901017383
- Primary Citation Related Structures:
1JFB, 1JFC - PubMed Abstract:
Crystal structures of the nitric oxide reductase cytochrome P450nor (P450nor) in the ferric resting and the ferrous carbonmonoxy (CO) states have been determined at 1.00 and 1.05 A resolution, respectively. P450nor consists of 403 amino-acid residues (46 kDa) and is one of the largest proteins refined to this resolution so far. The final models have conventional R factors of 10.2% (ferric resting) and 11.7% (ferrous CO), with mean coordinate errors of 0.028 (ferric resting) and 0.030 A (ferrous CO) as calculated from inversion of the full positional least-squares matrix. Owing to the atomic resolution, novel features are found in the refined structures. Firstly, two orientations of the haem are observed both in the ferric resting and the ferrous CO states. Secondly, a disordered water molecule bound to the haem iron is found in the ferric resting state. In addition, the accurate structures at atomic resolution enabled the examination of general stereochemical parameters that are commonly used in refinement cycles of protein structures.
- RIKEN Harima Institute/SPring-8, 1-1-1 Kouto, Mikazuki, Sayo, Hyogo 679-5148, Japan.
Organizational Affiliation: