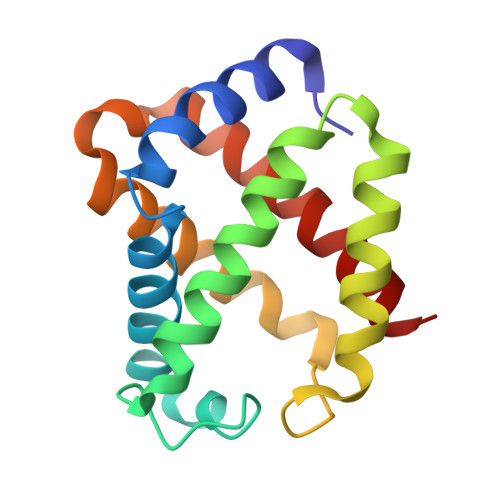
Crystal Structures of Unligated and CN-Ligated Glycera dibranchiata Monomer Ferric Hemoglobin Components III and IV
Park, H.J., Yang, C., Treff, N., Satterlee, J.D., Kang, C.(2002) Proteins 49: 49-60
- PubMed: 12211015
- DOI: https://doi.org/10.1002/prot.10199
- Primary Citation Related Structures:
1JF3, 1JF4, 1JL6, 1JL7 - PubMed Abstract:
Erythrocytes of the marine annelid, Glycera dibranchiata, contain a mixture of monomeric and polymeric hemoglobins. There are three major monomer hemoglobin components, II, III, IV (also called GMH2, 3, and 4), that have been highly purified and well characterized. We have now crystallized GMH3 and GMH4 and determined their structures to 1.4-1.8 A resolution. The structures were determined for these two monomer hemoglobins in the oxidized (Fe3+, ferric, or met-) forms in both the unligated and cyanide-ligated states. This work differs from two published, refined structures of a Glycera dibranchiata monomer hemoglobin, which has a sequence that is substantially different from any bona fide major monomer hemoglobins (GMH2, 3, or 4). The high-resolution crystal structures (presented here) and the previous NMR structure of CO-ligated GMH4, provide a basis for interpreting structure/function details of the monomer hemoglobins. These details include: (1) the strong correlation between temperature factor and NMR dynamics for respective protein forms; (2) the unique nature of the HisE7Leu primary sequence substitutions in GMH3 and GMH4 and their impact on cyanide ion binding kinetics; (3) the LeuB10Phe difference between GMH3 and GMH4 and its impact on ligand binding; and (4) elucidation of changes in the structural details of the distal and proximal heme pockets upon cyanide binding.
- School of Molecular Biosciences, Washington State University, Pullman, Washington 99164-4660, USA.
Organizational Affiliation: