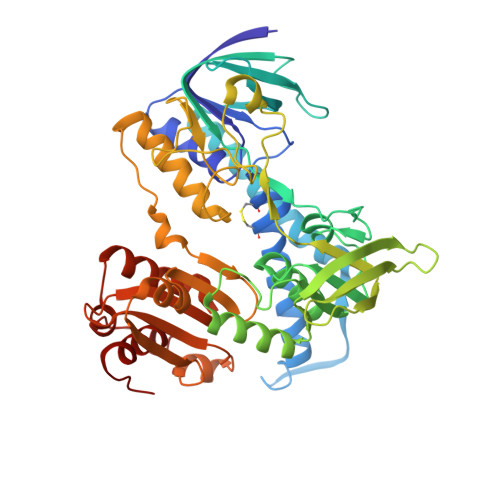
Crystal structure of eucaryotic E3, lipoamide dehydrogenase from yeast.
Toyoda, T., Suzuki, K., Sekiguchi, T., Reed, L.J., Takenaka, A.(1998) J Biochem 123: 668-674
- PubMed: 9538259 Search on PubMed
- DOI: https://doi.org/10.1093/oxfordjournals.jbchem.a021989
- Primary Citation Related Structures:
1JEH - PubMed Abstract:
The crystal structure of eucaryotic lipoamide dehydrogenase from yeast has been determined by an X-ray analysis at 2.7 (partially at 2.4) A resolution. The enzyme has two identical subunits related by a pseudo twofold symmetry. The tertiary structure is similar to those of other procaryotic enzymes. The active site, consisting of FAD, Cys44, and Cys49 from one subunit and His457' from the other subunit, is highly conserved. This enzyme is directly bound to the core protein E2 of the 2-oxoglutarate dehydrogenase complex, whereas it is bound to the pyruvate dehydrogenase complex through a protein X. The calculated electrostatic potential suggests two characteristic regions for binding with these two proteins.
- Department of Life Science, Faculty of Bioscience and Biotechnology, Tokyo Institute of Technology, Yokohama 226-8501.
Organizational Affiliation: